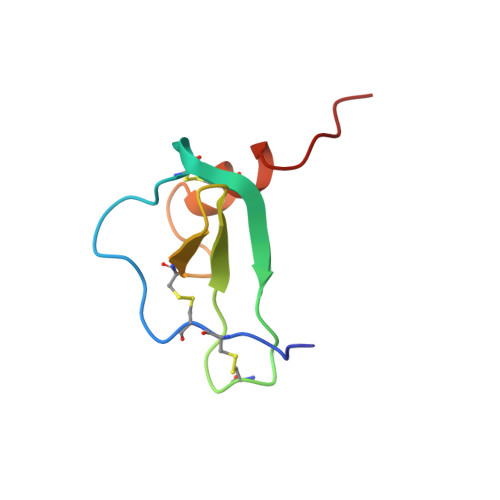
Solution structure of the human CC chemokine 2: A monomeric representative of the CC chemokine subtype.
Sticht, H., Escher, S.E., Schweimer, K., Forssmann, W.G., Rosch, P., Adermann, K.(1999) Biochemistry 38: 5995-6002
- PubMed: 10320325 Search on PubMed
- DOI: https://doi.org/10.1021/bi990065i
- Primary Citation Related Structures:
2HCC - PubMed Abstract:
HCC-2, a 66-amino acid residue human CC chemokine, was reported to induce chemotaxis on monocytes, T-lymphocytes, and eosinophils. The three-dimensional structure of HCC-2 has been determined by 1H nuclear magnetic resonance (NMR) spectroscopy and restrained molecular dynamics calculations on the basis of 871 experimental restraints. The structure is well-defined, exhibiting average root-mean-square deviations of 0.58 and 0.96 A for the backbone heavy atoms and all heavy atoms of residues 5-63, respectively. In contrast to most other chemokines, subtle structural differences impede dimer formation of HCC-2 in a concentration range of 0.1 microM to 2 mM. HCC-2, however, exhibits the same structural elements as the other chemokines, i.e., a triple-stranded antiparallel beta-sheet covered by an alpha-helix, showing that the chemokine fold is not influenced by quaternary interactions. Structural investigations with a HCC-2 mutant prove that a third additional disulfide bond present in wild-type HCC-2 is not necessary for maintaining the relative orientation of the helix and the beta-sheet.
- Lehrstuhl für Biopolymere, Universität Bayreuth, D-95440 Bayreuth, Germany. Heinrich.Sticht@Uni-Bayreuth.de
Organizational Affiliation: