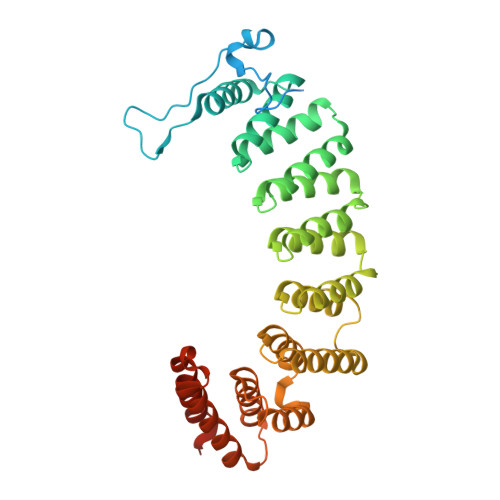
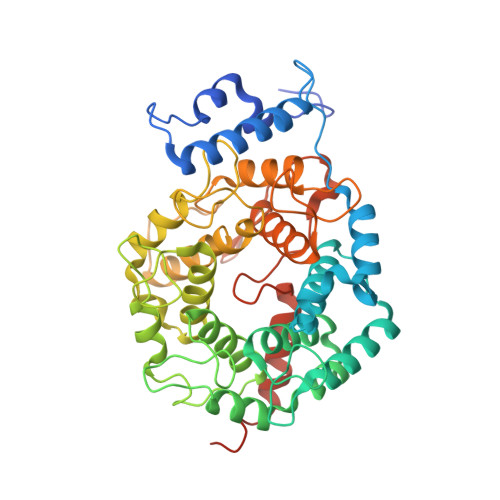
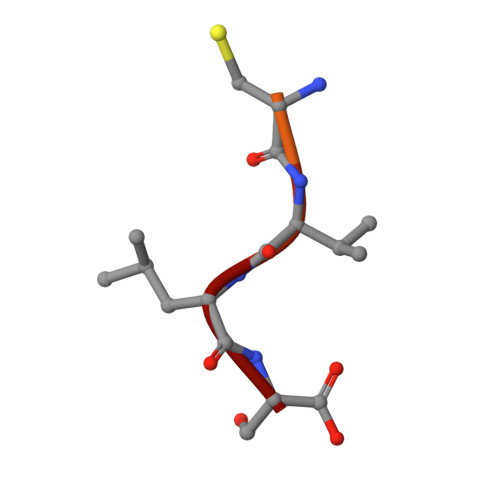
Conversion of protein farnesyltransferase to a geranylgeranyltransferase.
Terry, K.L., Casey, P.J., Beese, L.S.(2006) Biochemistry 45: 9746-9755
- PubMed: 16893176 Search on PubMed
- DOI: https://doi.org/10.1021/bi060295e
- Primary Citation Related Structures:
2H6F, 2H6G, 2H6H, 2H6I - PubMed Abstract:
Posttranslational modifications are essential for the proper function of a number of proteins in the cell. One such modification, the covalent attachment of a single isoprenoid lipid (prenylation), is carried out by the CaaX prenyltransferases, protein farnesyltransferase (FTase) and protein geranylgeranyltransferase type-I (GGTase-I). Substrate proteins of these two enzymes are involved in a variety of cellular functions but are largely associated with signal transduction. These modified proteins include members of the Ras superfamily, heterotrimeric G-proteins, centromeric proteins, and a number of proteins involved in nuclear integrity. Although FTase and GGTase-I are highly homologous, they are quite selective for their substrates, particularly for their isoprenoid diphosphate substrates, FPP and GGPP, respectively. Here, we present both crystallographic and kinetic analyses of mutants designed to explore this isoprenoid specificity and demonstrate that this specificity is dependent upon two enzyme residues in the beta subunits of the enzymes, W102beta and Y365beta in FTase (T49beta and F324beta, respectively, in GGTase-I).
- Department of Biochemistry, Duke University Medical Center, Durham, North Carolina 27710, USA.
Organizational Affiliation: