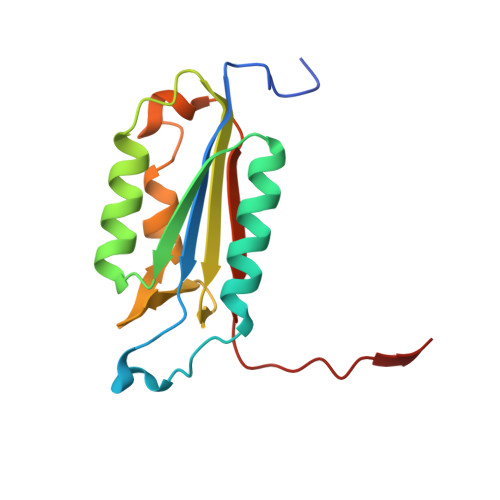
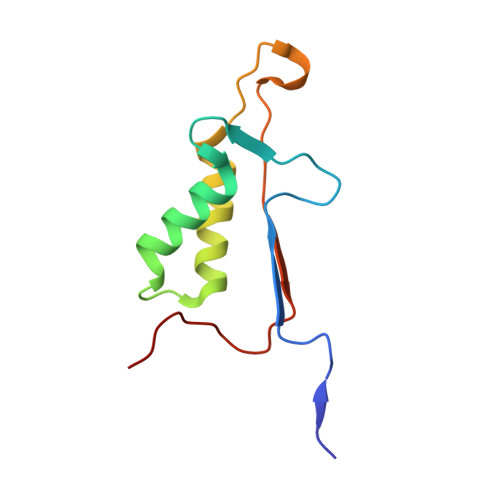
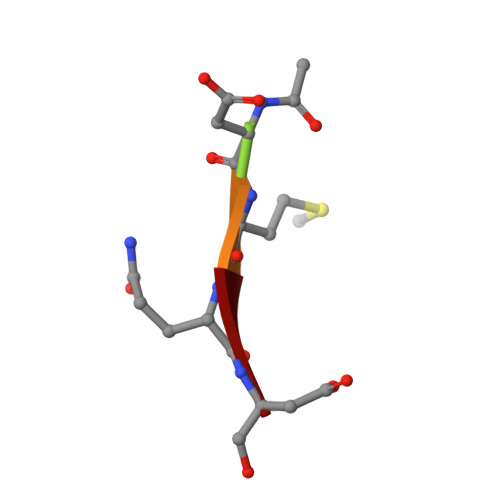
Structural and kinetic analysis of caspase-3 reveals role for s5 binding site in substrate recognition
Fang, B., Boross, P.I., Tozser, J., Weber, I.T.(2006) J Mol Biology 360: 654-666
- PubMed: 16781734 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.05.041
- Primary Citation Related Structures:
2H5I, 2H5J, 2H65 - PubMed Abstract:
The molecular basis for the substrate specificity of human caspase-3 has been investigated using peptide analog inhibitors and substrates that vary at the P2, P3, and P5 positions. Crystal structures were determined of caspase-3 complexes with the substrate analogs at resolutions of 1.7 A to 2.3 A. Differences in the interactions of caspase-3 with the analogs are consistent with the Ki values of 1.3 nM, 6.5 nM, and 12.4 nM for Ac-DEVD-Cho, Ac-VDVAD-Cho and Ac-DMQD-Cho, respectively, and relative kcat/Km values of 100%, 37% and 17% for the corresponding peptide substrates. The bound peptide analogs show very similar interactions for the main-chain atoms and the conserved P1 Asp and P4 Asp, while interactions vary for P2 and P3. P2 lies in a hydrophobic S2 groove, consistent with the weaker inhibition of Ac-DMQD-Cho with polar P2 Gln. S3 is a surface hydrophilic site with favorable polar interactions with P3 Glu in Ac-DEVD-Cho. Ac-DMQD-Cho and Ac-VDVAD-Cho have hydrophobic P3 residues that are not optimal in the polar S3 site, consistent with their weaker inhibition. A hydrophobic S5 site was identified for caspase-3, where the side-chains of Phe250 and Phe252 interact with P5 Val of Ac-VDVAD-Cho, and enclose the substrate-binding site by conformational change. The kinetic importance of hydrophobic P5 residues was confirmed by more efficient hydrolysis of caspase-3 substrates Ac-VDVAD-pNA and Ac-LDVAD-pNA compared with Ac-DVAD-pNA. In contrast, caspase-7 showed less efficient hydrolysis of the substrates with P5 Val or Leu compared with Ac-DVAD-pNA. Caspase-3 and caspase-2 share similar hydrophobic S5 sites, while caspases 1, 7, 8 and 9 do not have structurally equivalent hydrophobic residues; these caspases are likely to differ in their selectivity for the P5 position of substrates. The distinct selectivity for P5 will help define the particular substrates and signaling pathways associated with each caspase.
- Department of Biology, Molecular Basis of Disease, Georgia State University, Atlanta, GA 30303, USA.
Organizational Affiliation: