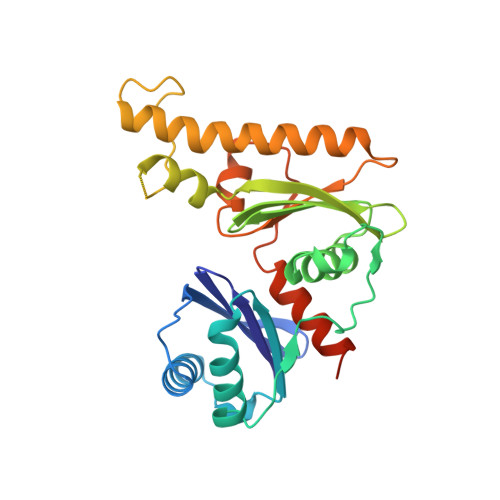
Structure of the Type III Pantothenate Kinase from Bacillus anthracis at 2.0 A Resolution: Implications for Coenzyme A-Dependent Redox Biology.
Nicely, N.I., Parsonage, D., Paige, C., Newton, G.L., Fahey, R.C., Leonardi, R., Jackowski, S., Mallett, T.C., Claiborne, A.(2007) Biochemistry 46: 3234-3245
- PubMed: 17323930 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi062299p
- Primary Citation Related Structures:
2H3G - PubMed Abstract:
Coenzyme A (CoASH) is the major low-molecular weight thiol in Staphylococcus aureus and a number of other bacteria; the crystal structure of the S. aureus coenzyme A-disulfide reductase (CoADR), which maintains the reduced intracellular state of CoASH, has recently been reported [Mallett, T.C., Wallen, J.R., Karplus, P.A., Sakai, H., Tsukihara, T., and Claiborne, A. (2006) Biochemistry 45, 11278-89]. In this report we demonstrate that CoASH is the major thiol in Bacillus anthracis; a bioinformatics analysis indicates that three of the four proteins responsible for the conversion of pantothenate (Pan) to CoASH in Escherichia coli are conserved in B. anthracis. In contrast, a novel type III pantothenate kinase (PanK) catalyzes the first committed step in the biosynthetic pathway in B. anthracis; unlike the E. coli type I PanK, this enzyme is not subject to feedback inhibition by CoASH. The crystal structure of B. anthracis PanK (BaPanK), solved using multiwavelength anomalous dispersion data and refined at a resolution of 2.0 A, demonstrates that BaPanK is a new member of the Acetate and Sugar Kinase/Hsc70/Actin (ASKHA) superfamily. The Pan and ATP substrates have been modeled into the active-site cleft; in addition to providing a clear rationale for the absence of CoASH inhibition, analysis of the Pan-binding pocket has led to the development of two new structure-based motifs (the PAN and INTERFACE motifs). Our analyses also suggest that the type III PanK in the spore-forming B. anthracis plays an essential role in the novel thiol/disulfide redox biology of this category A biodefense pathogen.
- Center for Structural Biology, Wake Forest University School of Medicine, Winston-Salem, North Carolina 27157, USA.
Organizational Affiliation: