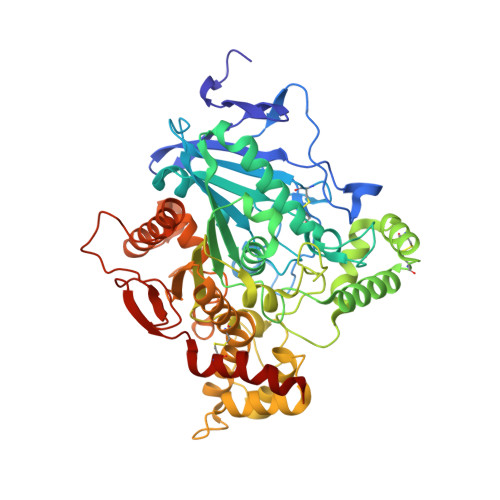
Crystal structures of acetylcholinesterase in complex with HI-6, Ortho-7 and obidoxime: Structural basis for differences in the ability to reactivate tabun conjugates.
Ekstrom, F., Pang, Y.P., Boman, M., Artursson, E., Akfur, C., Borjegren, S.(2006) Biochem Pharmacol 72: 597-607
- PubMed: 16876764 Search on PubMed
- DOI: https://doi.org/10.1016/j.bcp.2006.05.027
- Primary Citation Related Structures:
2GYU, 2GYV, 2GYW - PubMed Abstract:
Inhibition of acetylcholinesterase (AChE) by organophosphorus compounds (OPs) such as pesticides and nerve agents causes acute toxicity or death of the intoxicated individual. The inhibited AChE may be reactivated by certain oximes as antidotes for clinical treatment of OP-intoxications. Crystal structures of the oximes HI-6, Ortho-7 and obidoxime in complex with Mus musculus acetylcholinesterase (mAChE) reveal different roles of the peripheral anionic site (PAS) in the binding of the oximes. A limited structural change of the side chains of Trp286 and Asp74 facilitates the intercalation of the 4-carboxylamide pyridinium ring of HI-6 between the side chains of Tyr124 and Trp286. The 2-carboxyimino pyridinium ring of HI-6 is accommodated at the entrance of the catalytic site with the oximate forming a hydrogen bond to the main-chain nitrogen atom of Phe295. In contrast to HI-6, the coordination of Ortho-7 and obidoxime within the PAS is facilitated by an extended structural change of Trp286 that allows one of the carboxyimino pyridinium rings to form a cation-pi interaction with the aromatic groups of Tyr72 and Trp286. The central chain of Ortho-7 and obidoxime is loosely coordinated in the active-site gorge, whereas the second carboxyimino pyridinium ring is accommodated in the vicinity of the phenol ring of Tyr337. The structural data clearly show analogous coordination of Ortho-7 and obidoxime within the active-site gorge of AChE. Different ability to reactivate AChE inhibited by tabun is shown in end-point reactivation experiments where HI-6, Ortho-7 and obidoxime showed an efficiency of 1, 45 and 38%, respectively. The low efficiency of HI-6 and the significantly higher efficiency of Ortho-7 and obidoxime may be explained by the differential binding of the oximes in the PAS and active-site gorge of AChE.
- Swedish Defense Research Agency, Division of NBC Defense, S-901 82, Umeå, Sweden. fredrik.ekstrom@foi.se
Organizational Affiliation: