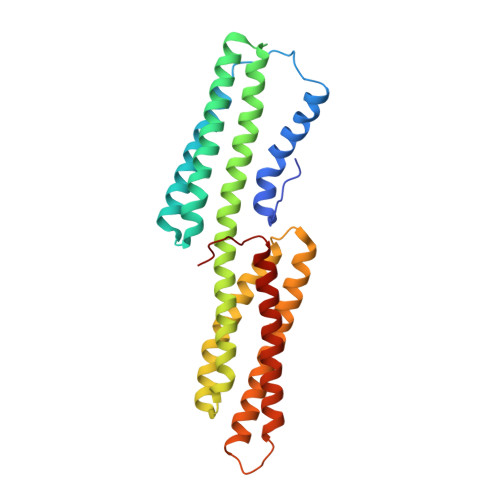
Shigella applies molecular mimicry to subvert vinculin and invade host cells.
Izard, T., Tran Van Nhieu, G., Bois, P.R.(2006) J Cell Biol 175: 465-475
- PubMed: 17088427 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1083/jcb.200605091
- Primary Citation Related Structures:
2GWW, 2HSQ - PubMed Abstract:
Shigella flexneri, the causative agent of bacillary dysentery, injects invasin proteins through a type III secretion apparatus upon contacting the host cell, which triggers pathogen internalization. The invasin IpaA is essential for S. flexneri pathogenesis and binds to the cytoskeletal protein vinculin to facilitate host cell entry. We report that IpaA harbors two vinculin-binding sites (VBSs) within its C-terminal domain that bind to and activate vinculin in a mutually exclusive fashion. Only the highest affinity C-terminal IpaA VBS is necessary for efficient entry and cell-cell spread of S. flexneri, whereas the lower affinity VBS appears to contribute to vinculin recruitment at entry foci of the pathogen. Finally, the crystal structures of vinculin in complex with the VBSs of IpaA reveal the mechanism by which IpaA subverts vinculin's functions, where S. flexneri utilizes a remarkable level of molecular mimicry of the talin-vinculin interaction to activate vinculin. Mimicry of vinculin's interactions may therefore be a general mechanism applied by pathogens to infect the host cell.
- Cell Adhesion Laboratory, Department of Oncology, St. Jude Children's Research Hospital, Memphis, TN 38105, USA. tina.izard@stjude.org
Organizational Affiliation: