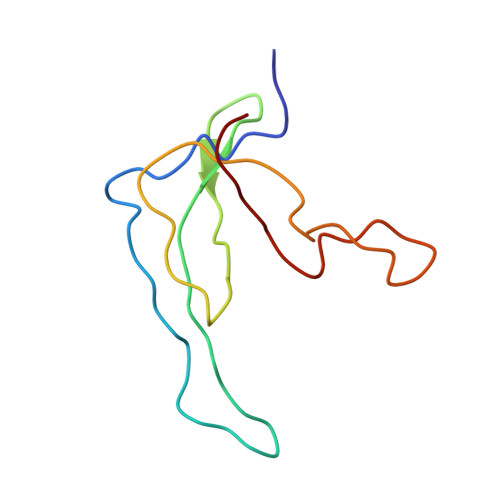
Refined structure of the gene 5 DNA binding protein from bacteriophage fd.
Brayer, G.D., McPherson, A.(1983) J Mol Biology 169: 565-596
- PubMed: 6684697 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(83)80065-5
- Primary Citation Related Structures:
2GN5 - PubMed Abstract:
The three-dimensional structure of the gene 5 DNA binding protein (G5BP) from bacteriophage fd has been determined from a combination of multiple isomorphous replacement techniques, partial refinements and deleted fragment difference Fourier syntheses. The structure was refined using restrained parameter least-squares and difference Fourier methods to a final residual of R = 0.217 for the 3528 statistically significant reflections present to 2.3 A resolution. In addition to the 682 atoms of the protein, 12 solvent molecules were included. We describe here the dispositions and orientations of the amino acid side-chains and their interactions as visualized in the G5BP structure. The G5BP monomer of 87 peptide units is almost entirely in the beta-conformation, organized as a three-stranded sheet, a two-stranded beta-ribbon and a broad connecting loop. There is no alpha-helix present in the molecule. Two G5BP monomers are tightly interlocked about an intermolecular dyad axis to form a compact dimer unit of about 55 A X 45 A X 36 A. The dimer is characterized by two symmetry-related antiparallel clefts that traverse the monomer surfaces essentially perpendicular to the dyad axis. From the three-stranded antiparallel beta-sheet, formed from the first two-thirds of the sequence, extend three tyrosine residues (26, 34, 41), a lysine (46) and two arginine residues (16, 21) that, as indicated by other physical and chemical experiments, are directly involved in DNA binding. Other residues likely to share binding responsibility are arginine 80 extending from the beta-ribbon and phenylalanine 73 from the tip of this loop, but as provided, however, by the opposite monomer within each G5BP dimer pair. Thus, both symmetry-related DNA binding sites have a composite nature and include contributions from both elements of the dimer. The gene 5 dimer is clearly the active binding species, and the two monomers within the dyad-related pair are so structurally contiguous that one cannot be certain whether the isolated monomer would maintain its observed crystal structure. This linkage is manifested primarily as a skeletal core of hydrophobic residues that extends from the center of each monomer continuously through an intermolecular beta-barrel that joins the pair. Protruding from the major area of density of each monomer is an elongated wing of tenuous structure comprising residues 15 through 32, which is, we believe, intimately involved in DNA binding. This wing appears to be dynamic and mobile, even in the crystal and, therefore, is likely to undergo conformational change in the presence of the ligand.