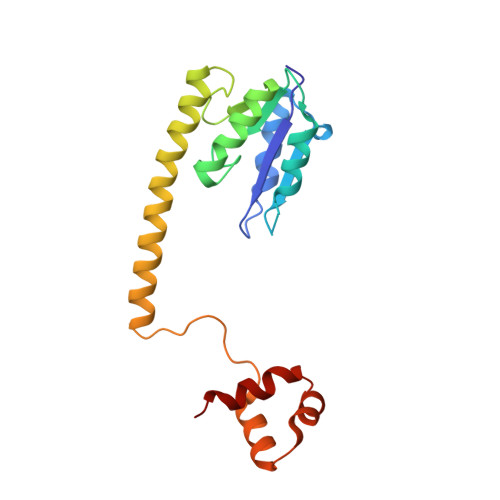
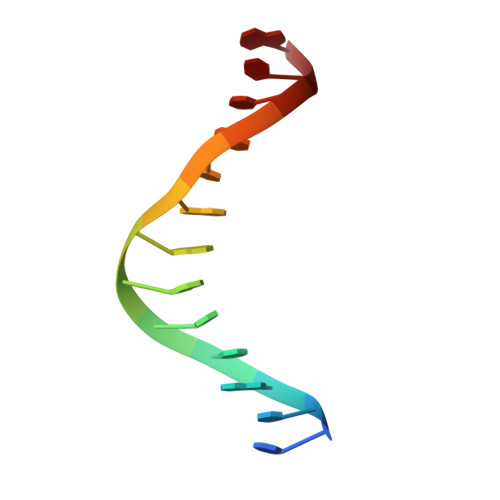
Implications of structures of synaptic tetramers of gamma delta resolvase for the mechanism of recombination.
Kamtekar, S., Ho, R.S., Cocco, M.J., Li, W., Wenwieser, S.V., Boocock, M.R., Grindley, N.D., Steitz, T.A.(2006) Proc Natl Acad Sci U S A 103: 10642-10647
- PubMed: 16807292 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0604062103
- Primary Citation Related Structures:
2GM4, 2GM5 - PubMed Abstract:
The structures of two mutants of the site-specific recombinase, gammadelta resolvase, that form activated tetramers have been determined. One, at 3.5-A resolution, forms a synaptic intermediate of resolvase that is covalently linked to two cleaved DNAs, whereas the other is of an unliganded structure determined at 2.1-A resolution. Comparisons of the four known tetrameric resolvase structures show that the subunits interact through the formation of a common core of four helices. The N-terminal halves of these helices superimpose well on each other, whereas the orientations of their C termini are more variable. The catalytic domains of resolvase in the unliganded structure are arranged asymmetrically, demonstrating that their positions can move substantially while preserving the four-helix core that forms the tetramer. These results suggest that the precleavage synaptic tetramer of gammadelta resolvase, whose structure is not known, may be formed by a similar four-helix core, but differ in the relative orientations of its catalytic and DNA-binding domains.
- Departments of Molecular Biophysics, and Howard Hughes Medical Institute, Yale University, New Haven, CT 06520-8114, USA.
Organizational Affiliation: