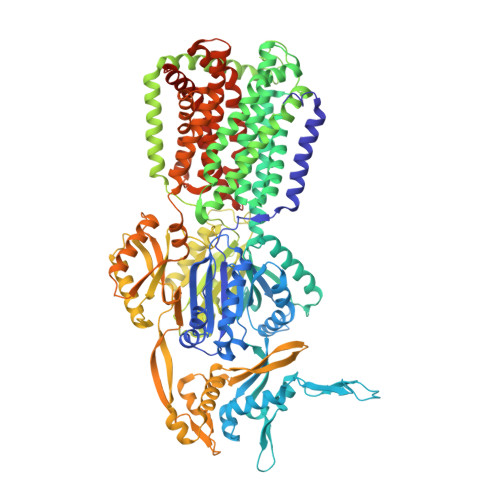
Structural Asymmetry of AcrB Trimer Suggests a Peristaltic Pump Mechanism.
Seeger, M.A., Schiefner, A., Eicher, T., Verrey, F., Diederichs, K., Pos, K.M.(2006) Science 313: 1295-1298
- PubMed: 16946072 Search on PubMed
- DOI: https://doi.org/10.1126/science.1131542
- Primary Citation Related Structures:
2GIF, 2HRT - PubMed Abstract:
The AcrA/AcrB/TolC complex spans the inner and outer membranes of Escherichia coli and serves as its major drug-resistance pump. Driven by the proton motive force, it mediates the efflux of bile salts, detergents, organic solvents, and many structurally unrelated antibiotics. Here, we report a crystallographic structure of trimeric AcrB determined at 2.9 and 3.0 angstrom resolution in space groups that allow asymmetry of the monomers. This structure reveals three different monomer conformations representing consecutive states in a transport cycle. The structural data imply an alternating access mechanism and a novel peristaltic mode of drug transport by this type of transporter.
- Institute of Physiology and Zurich Centre for Integrative Human Physiology (ZIHP), University of Zurich, Winterthurerstrasse 190, Zürich, Switzerland.
Organizational Affiliation: