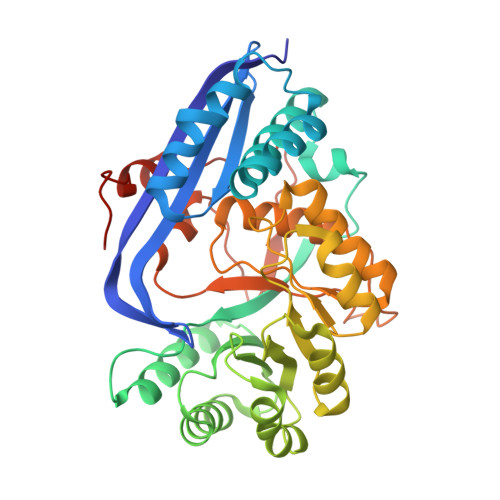
Crystal Structure of Mandelate Racemase/Muconate Lactonizing Enzyme from Bacillus Subtilis complexed with MG++ at 1.8 A
Malashkevich, V.N., Sauder, J.M., Schwinn, K.D., Emtage, S., Thompson, D.A., Rutter, M.E., Dickey, M., Groshong, C., Bain, K.T., Adams, J.M., Reyes, C., Rooney, I., Powell, A., Boice, A., Gheyi, T., Ozyurt, S., Atwell, S., Wasserman, S.R., Burley, S.K., Sali, A., Babbitt, P., Pieper, U., Gerlt, J.A., Almo, S.C.To be published.