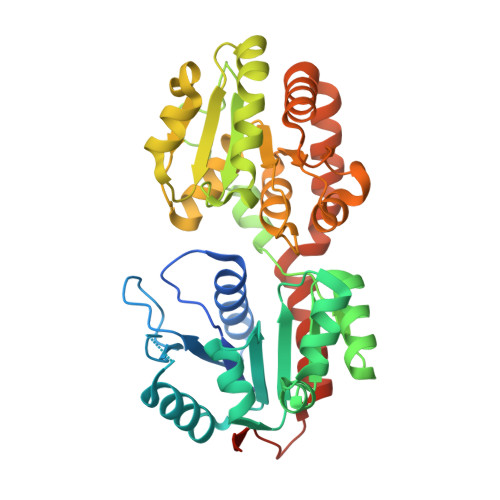
Molecular recognition and interfacial catalysis by the essential phosphatidylinositol mannosyltransferase PimA from mycobacteria.
Guerin, M.E., Kordulakova, J., Schaeffer, F., Svetlikova, Z., Buschiazzo, A., Giganti, D., Gicquel, B., Mikusova, K., Jackson, M., Alzari, P.M.(2007) J Biological Chem 282: 20705-20714
- PubMed: 17510062 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M702087200
- Primary Citation Related Structures:
2GEJ, 2GEK - PubMed Abstract:
Mycobacterial phosphatidylinositol mannosides (PIMs) and metabolically derived cell wall lipoglycans play important roles in host-pathogen interactions, but their biosynthetic pathways are poorly understood. Here we focus on Mycobacterium smegmatis PimA, an essential enzyme responsible for the initial mannosylation of phosphatidylinositol. The structure of PimA in complex with GDP-mannose shows the two-domain organization and the catalytic machinery typical of GT-B glycosyltransferases. PimA is an amphitrophic enzyme that binds mono-disperse phosphatidylinositol, but its transferase activity is stimulated by high concentrations of non-substrate anionic surfactants, indicating that the early stages of PIM biosynthesis involve lipid-water interfacial catalysis. Based on structural, calorimetric, and mutagenesis studies, we propose a model wherein PimA attaches to the membrane through its N-terminal domain, and this association leads to enzyme activation. Our results reveal a novel mode of phosphatidylinositol recognition and provide a template for the development of potential antimycobacterial compounds.
- Unité de Biochimie Structurale (CNRS URA 2185), Institut Pasteur, 25 rue du Docteur Roux, 75724 Paris Cedex 15, France.
Organizational Affiliation: