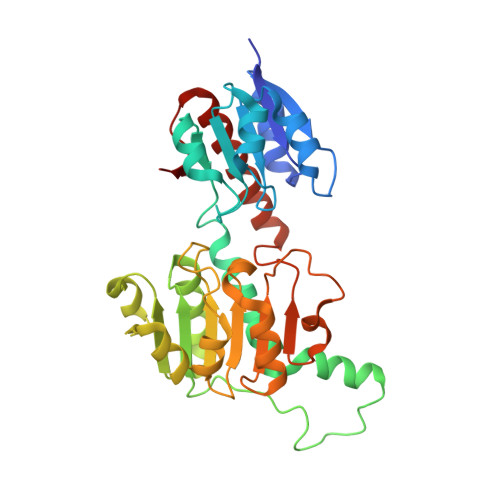
Structural basis of substrate specificity in human glyoxylate reductase/hydroxypyruvate reductase
Booth, M.P.S., Conners, R., Rumsby, G., Brady, R.L.(2006) J Mol Biology 360: 178-189
- PubMed: 16756993 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.05.018
- Primary Citation Related Structures:
2GCG, 2WWR - PubMed Abstract:
Human glyoxylate reductase/hydroxypyruvate reductase (GRHPR) is a D-2-hydroxy-acid dehydrogenase that plays a critical role in the removal of the metabolic by-product glyoxylate from within the liver. Deficiency of this enzyme is the underlying cause of primary hyperoxaluria type 2 (PH2) and leads to increased urinary oxalate levels, formation of kidney stones and renal failure. Here we describe the crystal structure of human GRHPR at 2.2 A resolution. There are four copies of GRHPR in the crystallographic asymmetric unit: in each homodimer, one subunit forms a ternary (enzyme+NADPH+reduced substrate) complex, and the other a binary (enzyme+NADPH) form. The spatial arrangement of the two enzyme domains is the same in binary and ternary forms. This first crystal structure of a true ternary complex of an enzyme from this family demonstrates the relationship of substrate and catalytic residues within the active site, confirming earlier proposals of the mode of substrate binding, stereospecificity and likely catalytic mechanism for these enzymes. GRHPR has an unusual substrate specificity, preferring glyoxylate and hydroxypyruvate, but not pyruvate. A tryptophan residue (Trp141) from the neighbouring subunit of the dimer is projected into the active site region and appears to contribute to the selectivity for hydroxypyruvate. This first crystal structure of a human GRHPR enzyme also explains the deleterious effects of naturally occurring missense mutations of this enzyme that lead to PH2.
- Department of Biochemistry, University of Bristol, Bristol BS8 1TD, UK.
Organizational Affiliation: