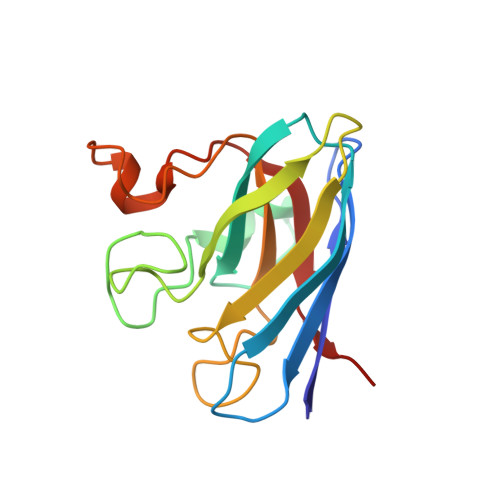
The Coupling between Disulphide Status, Metallation and Dimer Interface Strength in Cu/Zn Superoxide Dismutase
Hornberg, A., Logan, D.T., Marklund, S.L., Oliveberg, M.(2007) J Mol Biology 365: 333-342
- PubMed: 17070542 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.09.048
- Primary Citation Related Structures:
2GBT, 2GBU, 2GBV - PubMed Abstract:
The gain of neurotoxic function in amyotrophic lateral sclerosis (ALS) has been linked to misfolding of the homodimeric enzyme Cu/Zn superoxide dismutase (SOD). Here, we present the crystal structure of fully cysteine-depleted human SOD (SOD(CallA)), representing a reduced, marginally stable intermediate on the folding pathway in vivo that has also been implicated as neurotoxic precursor state. A hallmark of this species is that it fails to dimerize and becomes trapped as a monomer in the absence of the active-site metals. The crystallographic data show that removal of the C57-C146 disulphide bond sets free the interface loop IV in the apo protein, whereas the same loop remains unaffected in the holo protein. Thus, the low dimerisation propensity of disulphide-reduced apoSOD seems to be of entropic origin due to increased loop flexibility in the monomeric state: in the disulphide-reduced holo protein this gain in configurational entropy upon splitting of the dimer interface is reduced by the metal coordination.
- Department of Biochemistry, Umeå University, SE-901 87 Umeå, Sweden.
Organizational Affiliation: