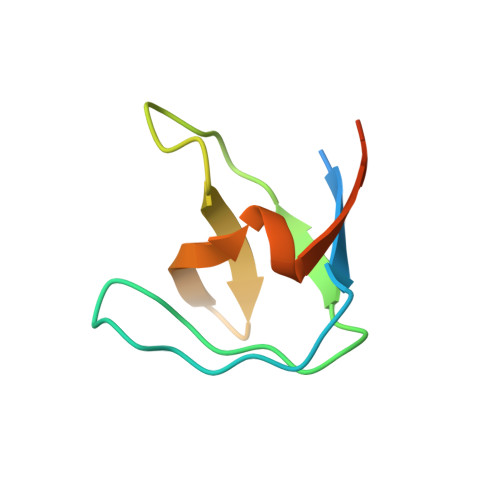
Solution structure of the Grb2 N-terminal SH3 domain complexed with a ten-residue peptide derived from SOS: direct refinement against NOEs, J-couplings and 1H and 13C chemical shifts.
Wittekind, M., Mapelli, C., Lee, V., Goldfarb, V., Friedrichs, M.S., Meyers, C.A., Mueller, L.(1997) J Mol Biology 267: 933-952
- PubMed: 9135122
- DOI: https://doi.org/10.1006/jmbi.1996.0886
- Primary Citation Related Structures:
1GBQ, 2GBQ, 3GBQ, 4GBQ - PubMed Abstract:
Refined ensembles of solution structures have been calculated for the N-terminal SH3 domain of Grb2 (N-SH3) complexed with the ac-VPPPVPPRRR-nh2 peptide derived from residues 1135 to 1144 of the mouse SOS-1 sequence. NMR spectra obtained from different combinations of both 13C-15N-labeled and unlabeled N-SH3 and SOS peptide fragment were used to obtain stereo-assignments for pro-chiral groups of the peptide, angle restraints via heteronuclear coupling constants, and complete 1H, 13C, and 15N resonance assignments for both molecules. One ensemble of structures was calculated using conventional methods while a second ensemble was generated by including additional direct refinements against both 1H and 13C(alpha)/13C(beta) chemical shifts. In both ensembles, the protein:peptide interface is highly resolved, reflecting the inclusion of 110 inter-molecular nuclear Overhauser enhancement (NOE) distance restraints. The first and second peptide-binding sub-sites of N-SH3 interact with structurally well-defined portions of the peptide. These interactions include hydrogen bonds and extensive hydrophobic contacts. In the third highly acidic sub-site, the conformation of the peptide Arg8 side-chain is partially ordered by a set of NOE restraints to the Trp36 ring protons. Overall, several lines of evidence point to dynamical averaging of peptide and N-SH3 side-chain conformations in the third subsite. These conformations are characterized by transient charge stabilized hydrogen bond interactions between the peptide arginine side-chain hydrogen bond donors and either single, or possibly multiple, acceptor(s) in the third peptide-binding sub-site.
- Macromolecular NMR Department, Bristol-Myers Squibb Pharmaceutical Research Institute, Princeton, NJ 08543-4000, USA.
Organizational Affiliation: