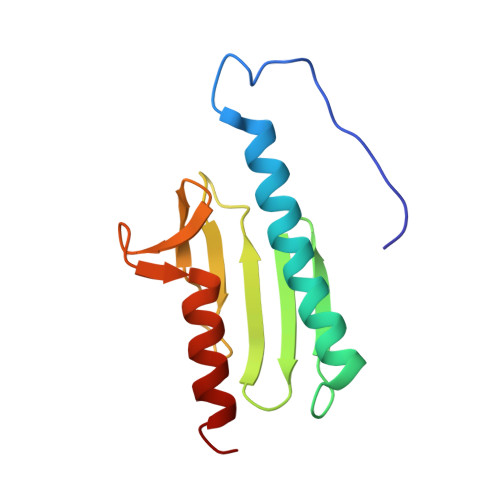
Yeast Frataxin Solution Structure, Iron Binding and Ferrochelatase Interaction
He, Y., Alam, S.L., Proteasa, S.V., Zhang, Y., Lesuisse, E., Dancid, E., Stemmler, T.L.(2004) Biochemistry 43: 16254-16262
- PubMed: 15610019
- DOI: https://doi.org/10.1021/bi0488193
- Primary Citation of Related Structures:
2GA5 - PubMed Abstract:
The mitochondrial protein frataxin is essential for cellular regulation of iron homeostasis. Although the exact function of frataxin is not yet clear, recent reports indicate the protein binds iron and can act as a mitochondrial iron chaperone to transport Fe(II) to ferrochelatase and ISU proteins within the heme and iron-sulfur cluster biosynthetic pathways, respectively. We have determined the solution structure of apo yeast frataxin to provide a structural basis of how frataxin binds and donates iron to the ferrochelatase. While the protein's alpha-beta-sandwich structural motif is similar to that observed for human and bacterial frataxins, the yeast structure presented in this report includes the full N-terminus observed for the mature processed protein found within the mitochondrion. In addition, NMR spectroscopy was used to identify frataxin amino acids that are perturbed by the presence of iron. Conserved acidic residues in the helix 1-strand 1 protein region undergo amide chemical shift changes in the presence of Fe(II), indicating a possible iron-binding site on frataxin. NMR spectroscopy was further used to identify the intermolecular binding interface between ferrochelatase and frataxin. Ferrochelatase appears to bind to frataxin's helical plane in a manner that includes its iron-binding interface.
- Department of Biochemistry and Molecular Biology, Wayne State University School of Medicine, Detroit, Michigan 48201, USA.
Organizational Affiliation: