Crystal Structure of the Bowman-Birk Inhibitor from Vigna unguiculata Seeds in Complex with {beta}-Trypsin at 1.55 A Resolution and Its Structural Properties in Association with Proteinases
Barbosa, J.A.R.G., Silva, L.P., Teles, R.C.L., Esteves, G.F., Azevedo, R.B., Ventura, M.M., Freitas, S.M.(2007) Biophys J 92: 1638-1650
- PubMed: 17142290 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1529/biophysj.106.090555
- Primary Citation Related Structures:
2G81 - PubMed Abstract:
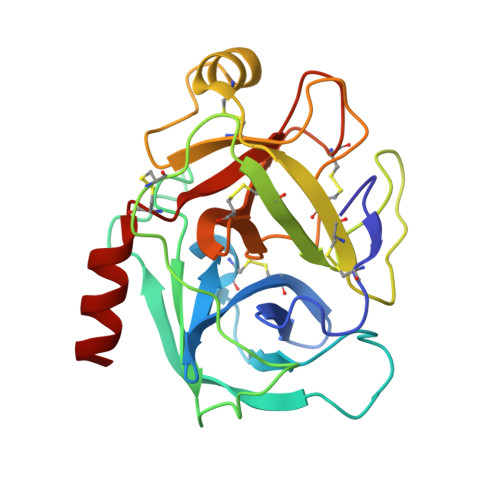
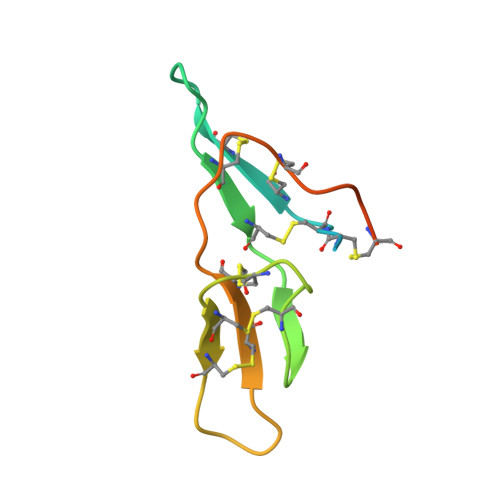
The structure of the Bowman-Birk inhibitor from Vigna unguiculata seeds (BTCI) in complex with beta-trypsin was solved and refined at 1.55 A to a crystallographic R(factor) of 0.154 and R(free) of 0.169, and represents the highest resolution for a Bowman-Birk inhibitor structure to date. The BTCI-trypsin interface is stabilized by hydrophobic contacts and hydrogen bonds, involving two waters and a polyethylene glycol molecule. The conformational rigidity of the reactive loop is characteristic of the specificity against trypsin, while hydrophobicity and conformational mobility of the antichymotryptic subdomain confer the self-association tendency, indicated by atomic force microscopy, of BTCI in complex and free form. When BTCI is in binary complexes, no significant differences in inhibition constants for producing a ternary complex with trypsin and chymotrypsin were detected. These results indicate that binary complexes present no conformational change in their reactive site for both enzymes confirming that these sites are structurally independent. The free chymotrypsin observed in the atomic force microscopy assays, when the ternary complex is obtained from BTCI-trypsin binary complex and chymotrypsin, could be related more to the self-association tendency between chymotrypsin molecules and the flexibility of the reactive site for this enzyme than to binding-related conformational changes.
- Centro de Biologia Molecular Estrutural, Laboratório Nacional de Luz Síncrotron, Campinas, Brazil.
Organizational Affiliation: