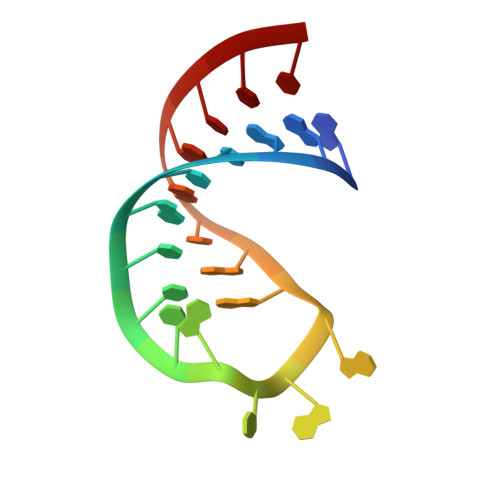
The testis-specific human protein RBMY recognizes RNA through a novel mode of interaction.
Skrisovska, L., Bourgeois, C.F., Stefl, R., Grellscheid, S.N., Kister, L., Wenter, P., Elliott, D.J., Stevenin, J., Allain, F.H.(2007) EMBO Rep 8: 372-379
- PubMed: 17318228 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.embor.7400910
- Primary Citation Related Structures:
2FY1 - PubMed Abstract:
The RBMY (RNA-binding motif gene on Y chromosome) protein encoded by the human Y chromosome is important for normal sperm development. Although its precise molecular RNA targets are unknown at present, it is suggested that human RBMY (hRBMY) participates in splicing in the testis. Using systematic evolution of ligands by exponential enrichment, we found that RNA stem-loops capped by a C(A)/(U)CAA pentaloop are high-affinity binding targets for hRBMY. Subsequent nuclear magnetic resonance structural determination of the hRBMY RNA recognition motif (RRM) in complex with a high-affinity target showed two distinct modes of RNA recognition. First, the RRM beta-sheet surface binds to the RNA loop in a sequence-specific fashion. Second, the beta2-beta3 loop of the hRBMY inserts into the major groove of the RNA stem. The first binding mode might be conserved in the paralogous protein heterogeneous nuclear RNP G, whereas the second mode of binding is found only in hRBMY. This structural difference could be at the origin of the function of RBMY in spermatogenesis.
- Institute of Molecular Biology and Biophysics, ETH Zurich, Swiss Federal Institute of Technology, 8093 Zurich, Switzerland.
Organizational Affiliation: