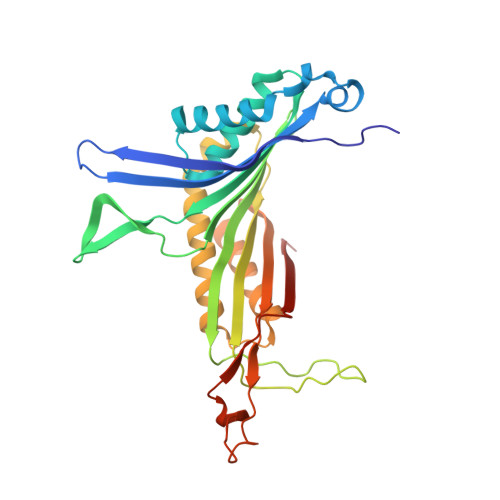
High pressure macromolecular crystallography: The 140-MPa crystal structure at 2.3 A resolution of urate oxidase, a 135-kDa tetrameric assembly
Colloc'h, N., Girard, E., Dhaussy, A.-C., Kahn, R., Ascone, I., Mezouar, M., Fourme, R.(2006) Biochim Biophys Acta 1764: 391-397
- PubMed: 16478683 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2006.01.006
- Primary Citation Related Structures:
2FUB - PubMed Abstract:
We report the three-dimensional structure determined by high-pressure macromolecular crystallography (HPMX) of a 135-kDa homo-tetrameric enzyme, urate oxidase from Aspergillus flavus complexed with its potent inhibitor 8-azaxanthin. Urate oxidase crystals are quite sensitive to pressure, as three-dimensional order is lost at about 180 MPa. A highly complete 2.3 A resolution data set was collected at 140 MPa, close to the critical pressure. Crystal structures at atmospheric pressure and at high pressure were refined in the orthorhombic space group I222 with final crystallographic R factors 14.1% and 16.1%, respectively. The effect of pressure on temperature factors, ordered water molecules, hydrogen bond lengths, contacts, buried surface areas as well as cavity volume was investigated. Results suggest that the onset of disruption of the tetrameric assembly by pressure has been captured in the crystalline state.
- Centre CYCERON, UMR 6185 Université de Caen-CNRS, Bd Becquerel, 14074 Caen, France.
Organizational Affiliation: