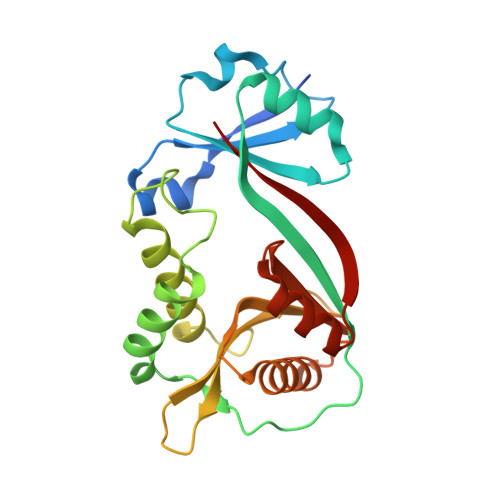
Crystal Structure of TDP-Fucosamine Acetyltransferase (WecD) from Escherichia coli, an Enzyme Required for Enterobacterial Common Antigen Synthesis.
Hung, M.N., Rangarajan, E., Munger, C., Nadeau, G., Sulea, T., Matte, A.(2006) J Bacteriol 188: 5606-5617
- PubMed: 16855251 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.00306-06
- Primary Citation Related Structures:
2FS5, 2FT0 - PubMed Abstract:
Enterobacterial common antigen (ECA) is a polysaccharide found on the outer membrane of virtually all gram-negative enteric bacteria and consists of three sugars, N-acetyl-d-glucosamine, N-acetyl-d-mannosaminuronic acid, and 4-acetamido-4,6-dideoxy-d-galactose, organized into trisaccharide repeating units having the sequence -->3)-alpha-d-Fuc4NAc-(1-->4)-beta-d-ManNAcA-(1-->4)-alpha-d-GlcNAc-(1-->. While the precise function of ECA is unknown, it has been linked to the resistance of Shiga-toxin-producing Escherichia coli (STEC) O157:H7 to organic acids and the resistance of Salmonella enterica to bile salts. The final step in the synthesis of 4-acetamido-4,6-dideoxy-d-galactose, the acetyl-coenzyme A (CoA)-dependent acetylation of the 4-amino group, is carried out by TDP-fucosamine acetyltransferase (WecD). We have determined the crystal structure of WecD in apo form at a 1.95-Angstrom resolution and bound to acetyl-CoA at a 1.66-Angstrom resolution. WecD is a dimeric enzyme, with each monomer adopting the GNAT N-acetyltransferase fold, common to a number of enzymes involved in acetylation of histones, aminoglycoside antibiotics, serotonin, and sugars. The crystal structure of WecD, however, represents the first structure of a GNAT family member that acts on nucleotide sugars. Based on this cocrystal structure, we have used flexible docking to generate a WecD-bound model of the acetyl-CoA-TDP-fucosamine tetrahedral intermediate, representing the structure during acetyl transfer. Our structural data show that WecD does not possess a residue that directly functions as a catalytic base, although Tyr208 is well positioned to function as a general acid by protonating the thiolate anion of coenzyme A.
- Biotechnology Research Institute, 6100 Royalmount Ave., Montreal QC, Canada H4P 2R2.
Organizational Affiliation: