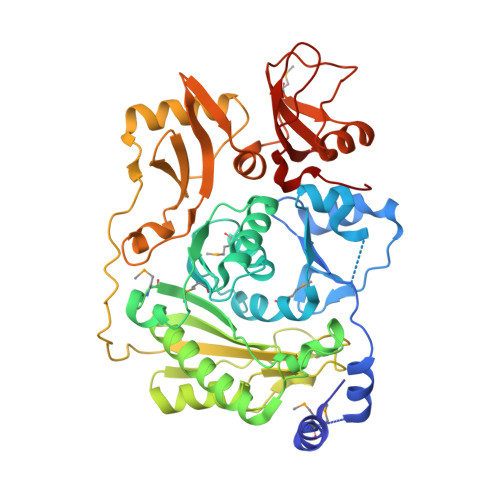
The structure of the RNA m5C methyltransferase YebU from Escherichia coli reveals a C-terminal RNA-recruiting PUA domain
Hallberg, B.M., Ericsson, U.B., Johnson, K.A., Andersen, N.M., Douthwaite, S., Nordlund, P., Beuscher IV, A.E., Erlandsen, H.(2006) J Mol Biology 360: 774-787
- PubMed: 16793063 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.05.047
- Primary Citation Related Structures:
2FRX - PubMed Abstract:
Nucleotide methylations are the most common type of rRNA modification in bacteria, and are introduced post-transcriptionally by a wide variety of site-specific enzymes. Three 5-methylcytidine (m(5)C) bases are found in the rRNAs of Escherichia coli and one of these, at nucleotide 1407 in 16 S rRNA, is the modification product of the methyltransferase (MTase) YebU (also called RsmF). YebU requires S-adenosyl-l-methionine (SAM) and methylates C1407 within assembled 30 S subunits, but not in naked 16 S rRNA or within tight-couple 70 S ribosomes. Here, we describe the three-dimensional structure of YebU determined by X-ray crystallography, and we present a molecular model for how YebU specifically recognizes, binds and methylates its ribosomal substrate. The YebU protein has an N-terminal SAM-binding catalytic domain with structural similarity to the equivalent domains in several other m(5)C RNA MTases including RsmB and PH1374. The C-terminal one-third of YebU contains a domain similar to that in pseudouridine synthases and archaeosine-specific transglycosylases (PUA-domain), which was not predicted by sequence alignments. Furthermore, YebU is predicted to contain extended regions of positive electrostatic potential that differ from other RNA-MTase structures, suggesting that YebU interacts with its RNA target in a different manner. Docking of YebU onto the 30 S subunit indicates that the PUA and MTase domains make several contacts with 16 S rRNA as well as with the ribosomal protein S12. The ribosomal protein interactions would explain why the assembled 30 S subunit, and not naked 16 S rRNA, is the preferred substrate for YebU.
- Department of Medical Biochemistry and Biophysics, Karolinska Institute, 17177 Stockholm, Sweden.
Organizational Affiliation: