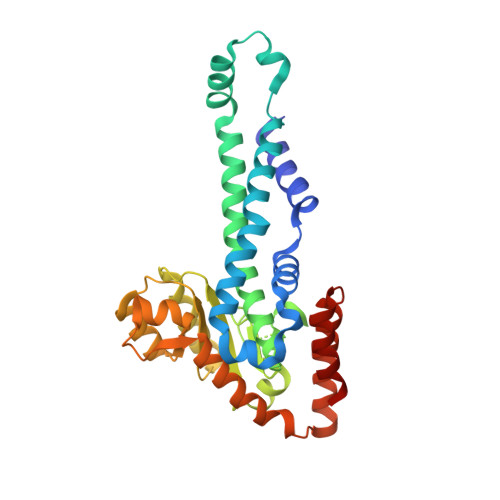
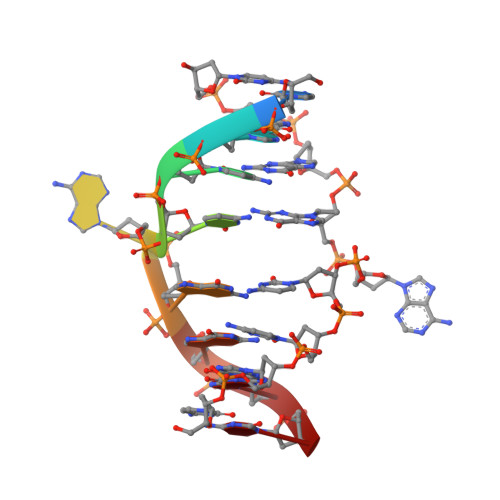
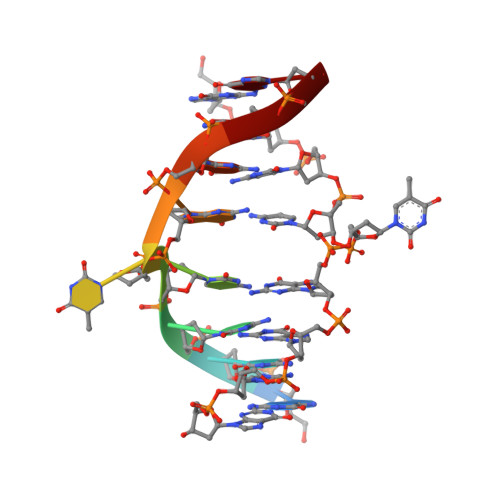
Nucleotide flips determine the specificity of the Ecl18kI restriction endonuclease
Bochtler, M., Szczepanowski, R.H., Tamulaitis, G., Grazulis, S., Czapinska, H., Manakova, E., Siksnys, V.(2006) EMBO J 25: 2219-2229
- PubMed: 16628220
- DOI: https://doi.org/10.1038/sj.emboj.7601096
- Primary Citation Related Structures:
2FQZ, 2GB7 - PubMed Abstract:
Restricion endonuclease Ecl18kI is specific for the sequence /CCNGG and cleaves it before the outer C to generate 5 nt 5'-overhangs. It has been suggested that Ecl18kI is evolutionarily related to NgoMIV, a 6-bp cutter that cleaves the sequence G/CCGGC and leaves 4 nt 5'-overhangs. Here, we report the crystal structure of the Ecl18kI-DNA complex at 1.7 A resolution and compare it with the known structure of the NgoMIV-DNA complex. We find that Ecl18kI flips both central nucleotides within the CCNGG sequence and buries the extruded bases in pockets within the protein. Nucleotide flipping disrupts Watson-Crick base pairing, induces a kink in the DNA and shifts the DNA register by 1 bp, making the distances between scissile phosphates in the Ecl18kI and NgoMIV cocrystal structures nearly identical. Therefore, the two enzymes can use a conserved DNA recognition module, yet recognize different sequences, and form superimposable dimers, yet generate different cleavage patterns. Hence, Ecl18kI is the first example of a restriction endonuclease that flips nucleotides to achieve specificity for its recognition site.
- International Institute of Molecular and Cell Biology, Warsaw, Poland. MBochtler@iimcb.gov.pl
Organizational Affiliation: