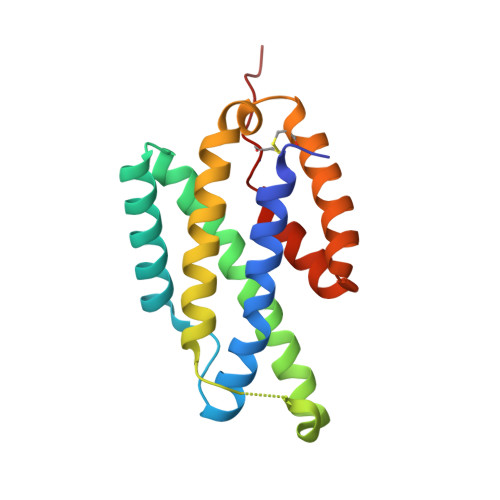
1.6A Crystal Structure of the Secreted Chorismate Mutase from Mycobacterium tuberculosis: Novel Fold Topology Revealed
Okvist, M., Dey, R., Sasso, S., Grahn, E., Kast, P., Krengel, U.(2006) J Mol Biology 357: 1483-1499
- PubMed: 16499927
- DOI: https://doi.org/10.1016/j.jmb.2006.01.069
- Primary Citation Related Structures:
2FP1, 2FP2 - PubMed Abstract:
The presence of exported chorismate mutases produced by certain organisms such as Mycobacterium tuberculosis has been shown to correlate with their pathogenicity. As such, these proteins comprise a new group of promising selective drug targets. Here, we report the high-resolution crystal structure of the secreted dimeric chorismate mutase from M. tuberculosis (*MtCM; encoded by Rv1885c), which represents the first 3D-structure of a member of this chorismate mutase family, termed the AroQ(gamma) subclass. Structures are presented both for the unliganded enzyme and for a complex with a transition state analog. The protomer fold resembles the structurally characterized (dimeric) Escherichia coli chorismate mutase domain, but exhibits a new topology, with helix H4 of *MtCM carrying the catalytic site residue missing in the shortened helix H1. Furthermore, the structure of each *MtCM protomer is significantly more compact and only harbors one active site pocket, which is formed entirely by one polypeptide chain. Apart from the structural model, we present evidence as to how the substrate may enter the active site.
- Department of Chemistry and Bioscience, Chalmers University of Technology, P.O. Box 462, SE-40530 Göteborg, Sweden.
Organizational Affiliation: