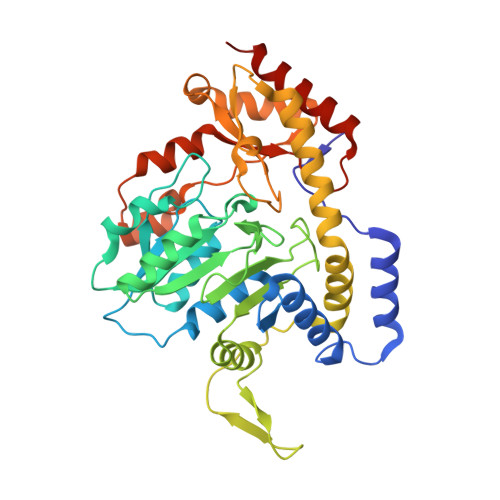
Structural and Functional Characterization of PseC, an Aminotransferase Involved in the Biosynthesis of Pseudaminic Acid, an Essential Flagellar Modification in Helicobacter pylori
Schoenhofen, I.C., Lunin, V.V., Julien, J.P., Li, Y., Ajamian, E., Matte, A., Cygler, M., Brisson, J.R., Aubry, A., Logan, S.M., Bhatia, S., Wakarchuk, W.W., Young, N.M.(2006) J Biological Chem 281: 8907-8916
- PubMed: 16421095 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M512987200
- Primary Citation Related Structures:
2FN6, 2FNI, 2FNU - PubMed Abstract:
Helicobacter pylori flagellin is heavily glycosylated with the novel sialic acid-like nonulosonate, pseudaminic acid (Pse). The glycosylation process is essential for assembly of functional flagellar filaments and consequent bacterial motility. Because motility is a key virulence factor for this and other important pathogens, the Pse biosynthetic pathway offers potential for novel therapeutic targets. From recent NMR analyses, we determined that the conversion of UDP-alpha-D-Glc-NAc to the central intermediate in the pathway, UDP-4-amino-4,6-dideoxy-beta-L-AltNAc, proceeds by formation of UDP-2-acetamido-2,6-dideoxy-beta-L-arabino-4-hexulose by the dehydratase/epimerase PseB (HP0840) followed with amino transfer by the aminotransferase, PseC (HP0366). The central role of PseC in the H. pylori Pse biosynthetic pathway prompted us to determine crystal structures of the native protein, its complexes with pyridoxal phosphate alone and in combination with the UDP-4-amino-4,6-dideoxy-beta-L-AltNAc product, the latter being converted to the external aldimine form in the active site of the enzyme. In the binding site, the AltNAc sugar ring adopts a 4C1 chair conformation, which is different from the predominant 1C4 form found in solution. The enzyme forms a homodimer where each monomer contributes to the active site, and these structures have permitted the identification of key residues involved in stabilization, and possibly catalysis, of the beta-L-arabino intermediate during the amino transfer reaction. The essential role of Lys183 in the catalytic event was confirmed by site-directed mutagenesis. This work presents for the first time a nucleotide-sugar aminotransferase co-crystallized with its natural ligand, and, in conjunction with the recent functional characterization of this enzyme, these results will assist in elucidating the aminotransferase reaction mechanism within the Pse biosynthetic pathway.
- Institute for Biological Sciences, National Research Council, Ottawa, Ontario K1A 0R6.
Organizational Affiliation: