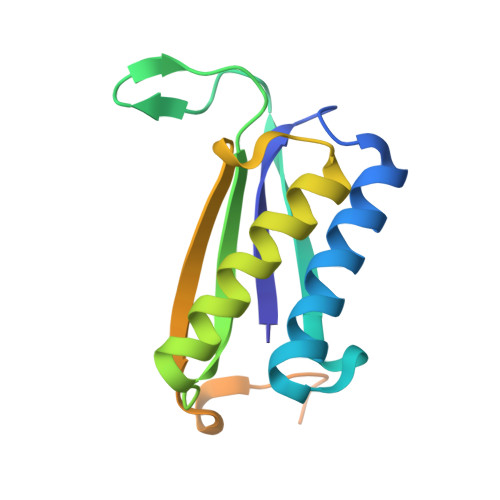
Crystal Structures of Native and Inactivated cis-3-Chloroacrylic Acid Dehalogenase: STRUCTURAL BASIS FOR SUBSTRATE SPECIFICITY AND INACTIVATION BY (R)-OXIRANE-2-CARBOXYLATE.
de Jong, R.M., Bazzacco, P., Poelarends, G.J., Johnson Jr., W.H., Kim, Y.J., Burks, E.A., Serrano, H., Thunnissen, A.M., Whitman, C.P., Dijkstra, B.W.(2007) J Biological Chem 282: 2440-2449
- PubMed: 17121835
- DOI: https://doi.org/10.1074/jbc.M608134200
- Primary Citation Related Structures:
2FLT, 2FLZ - PubMed Abstract:
The bacterial degradation pathways for the nematocide 1,3-dichloropropene rely on hydrolytic dehalogenation reactions catalyzed by cis- and trans-3-chloroacrylic acid dehalogenases (cis-CaaD and CaaD, respectively). X-ray crystal structures of native cis-CaaD and cis-CaaD inactivated by (R)-oxirane-2-carboxylate were elucidated. They locate four known catalytic residues (Pro-1, Arg-70, Arg-73, and Glu-114) and two previously unknown, potential catalytic residues (His-28 and Tyr-103'). The Y103F and H28A mutants of these latter two residues displayed reductions in cis-CaaD activity confirming their importance in catalysis. The structure of the inactivated enzyme shows covalent modification of the Pro-1 nitrogen atom by (R)-2-hydroxypropanoate at the C3 position. The interactions in the complex implicate Arg-70 or a water molecule bound to Arg-70 as the proton donor for the epoxide ring-opening reaction and Arg-73 and His-28 as primary binding contacts for the carboxylate group. This proposed binding mode places the (R)-enantiomer, but not the (S)-enantiomer, in position to covalently modify Pro-1. The absence of His-28 (or an equivalent) in CaaD could account for the fact that CaaD is not inactivated by either enantiomer. The cis-CaaD structures support a mechanism in which Glu-114 and Tyr-103' activate a water molecule for addition to C3 of the substrate and His-28, Arg-70, and Arg-73 interact with the C1 carboxylate group to assist in substrate binding and polarization. Pro-1 provides a proton at C2. The involvement of His-28 and Tyr-103' distinguishes the cis-CaaD mechanism from the otherwise parallel CaaD mechanism. The two mechanisms probably evolved independently as the result of an early gene duplication of a common ancestor.
- Laboratory of Biophysical Chemistry, Groningen Biomolecular Sciences and Biotechnology Institute, University of Groningen, Nijenborgh 4, 9747 AG Groningen, The Netherlands.
Organizational Affiliation: