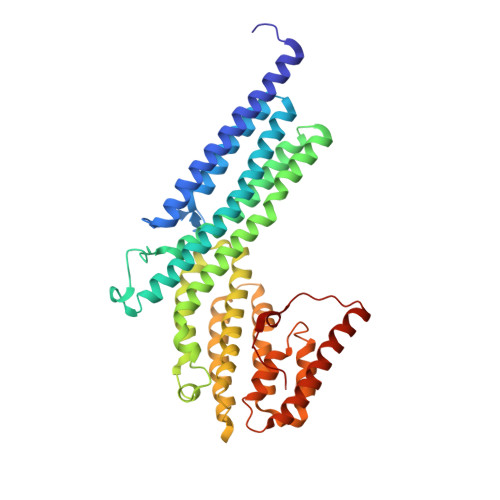
The structure of the exocyst subunit Sec6p defines a conserved architecture with diverse roles.
Sivaram, M.V., Furgason, M.L., Brewer, D.N., Munson, M.(2006) Nat Struct Mol Biol 13: 555-556
- PubMed: 16699513 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb1096
- Primary Citation Related Structures:
2FJI - PubMed Abstract:
The exocyst is a conserved protein complex essential for trafficking secretory vesicles to the plasma membrane. The structure of the C-terminal domain of the exocyst subunit Sec6p reveals multiple helical bundles, which are structurally and topologically similar to Exo70p and the C-terminal domains of Exo84p and Sec15, despite <10% sequence identity. The helical bundles appear to be evolutionarily related molecular scaffolds that have diverged to create functionally distinct exocyst proteins.
- Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical School, 364 Plantation Street, Worcester, Massachusetts 01605, USA.
Organizational Affiliation: