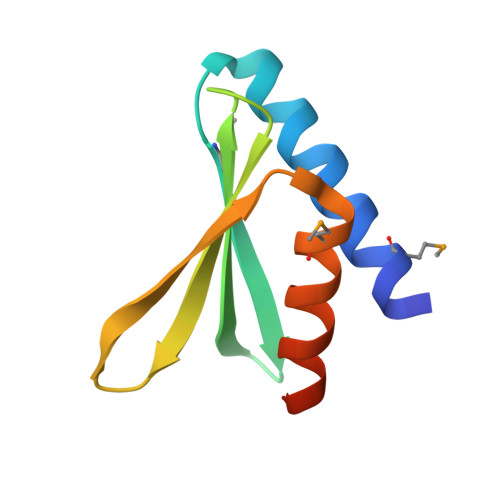
Novel x-ray structure of the YkuJ protein from Bacillus subtilis. Northeast Structural Genomics target SR360.
Kuzin, A.P., Abashidze, M., Forouhar, F., Vorobiev, S.M., Ho, C.K., Janjua, H., Cunningham, K., Conover, K., Ma, L.C., Xiao, R., Acton, T.B., Montelione, G.T., Tong, L., Hunt, J.F., Northeast Structural Genomics Consortium (NESG)To be published.