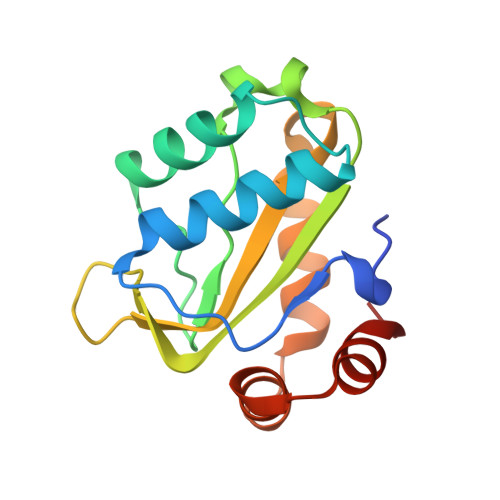
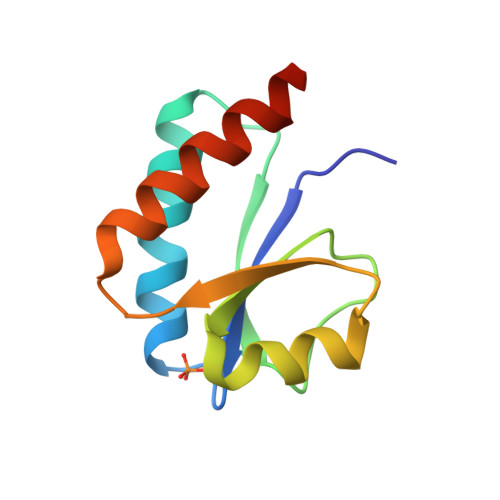
Solution Structure of a Post-transition State Analog of the Phosphotransfer Reaction between the A and B Cytoplasmic Domains of the Mannitol Transporter IIMannitol of the Escherichia coli Phosphotransferase System
Suh, J.Y., Cai, M., Williams Jr., D.C., Clore, G.M.(2006) J Biological Chem 281: 8939-8949
- PubMed: 16443929
- DOI: https://doi.org/10.1074/jbc.M513466200
- Primary Citation of Related Structures:
2FEW - PubMed Abstract:
The solution structure of the post-transition state complex between the isolated cytoplasmic A (IIAMtl) and phosphorylated B (phospho-IIBMtl) domains of the mannitol transporter of the Escherichia coli phosphotransferase system has been solved by NMR. The active site His-554 of IIAMtl was mutated to glutamine to block phosphoryl transfer activity, and the active site Cys-384 of IIBMtl (residues of IIBMtl are denoted in italic type) was substituted by serine to permit the formation of a stable phosphorylated form of IIBMtl. The two complementary interaction surfaces are predominantly hydrophobic, and two methionines on IIBMtl, Met-388 and Met-393, serve as anchors by interacting with two deep pockets on the surface of IIAMtl. With the exception of a salt bridge between the conserved Arg-538 of IIAMtl and the phosphoryl group of phospho-IIBMtl, electrostatic interactions between the two proteins are limited to the outer edges of the interface, are few in number, and appear to be weak. This accounts for the low affinity of the complex (Kd approximately 3.7 mm), which is optimally tuned to the intact biological system in which the A and B domains are expressed as a single polypeptide connected by a flexible 21-residue linker. The phosphoryl transition state can readily be modeled with no change in protein-protein orientation and minimal perturbations in both the backbone immediately adjacent to His-554 and Cys-384 and the side chains in close proximity to the phosphoryl group. Comparison with the previously solved structure of the IIAMtl-HPr complex reveals how IIAMtl uses the same interaction surface to recognize two structurally unrelated proteins and explains the much higher affinity of IIAMtl for HPr than IIBMtl.
- Laboratory of Chemical Physics, NIDDK, National Institutes of Health, Bethesda, Maryland 20892, USA.
Organizational Affiliation: