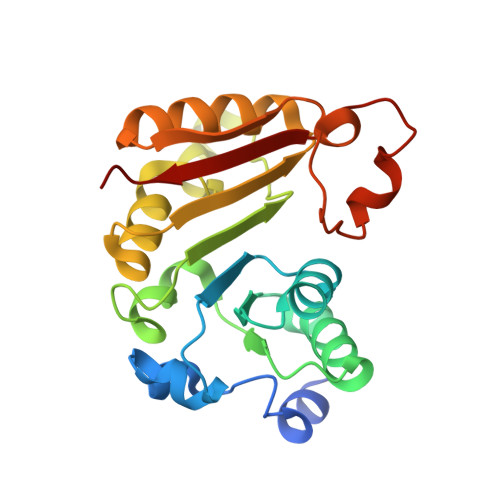
Crystal structure of Bacillus subtilis TrmB, the tRNA (m7G46) methyltransferase.
Zegers, I., Gigot, D., van Vliet, F., Tricot, C., Aymerich, S., Bujnicki, J.M., Kosinski, J., Droogmans, L.(2006) Nucleic Acids Res 34: 1925-1934
- PubMed: 16600901 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkl116
- Primary Citation Related Structures:
2FCA - PubMed Abstract:
The structure of Bacillus subtilis TrmB (BsTrmB), the tRNA (m7G46) methyltransferase, was determined at a resolution of 2.1 A. This is the first structure of a member of the TrmB family to be determined by X-ray crystallography. It reveals a unique variant of the Rossmann-fold methyltransferase (RFM) structure, with the N-terminal helix folded on the opposite site of the catalytic domain. The architecture of the active site and a computational docking model of BsTrmB in complex with the methyl group donor S-adenosyl-L-methionine and the tRNA substrate provide an explanation for results from mutagenesis studies of an orthologous enzyme from Escherichia coli (EcTrmB). However, unlike EcTrmB, BsTrmB is shown here to be dimeric both in the crystal and in solution. The dimer interface has a hydrophobic core and buries a potassium ion and five water molecules. The evolutionary analysis of the putative interface residues in the TrmB family suggests that homodimerization may be a specific feature of TrmBs from Bacilli, which may represent an early stage of evolution to an obligatory dimer.
- Laboratorium Ultrastructuur, Vrije Universiteit Brussel, Pleinlaan 2, B-1050 Brussel, Belgium.
Organizational Affiliation: