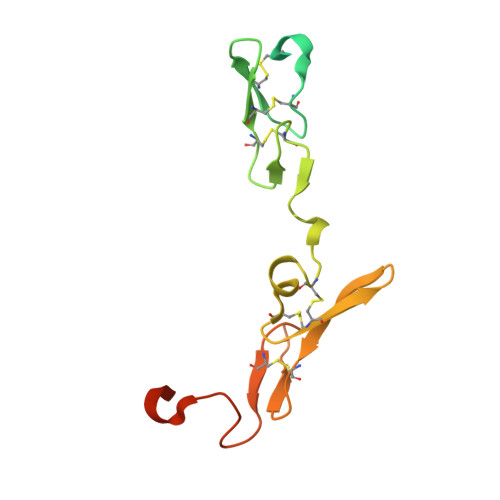
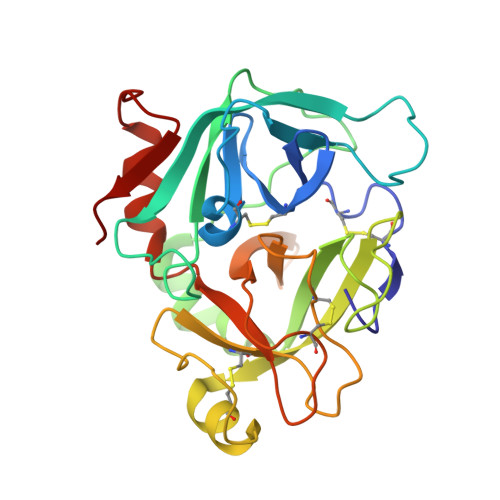
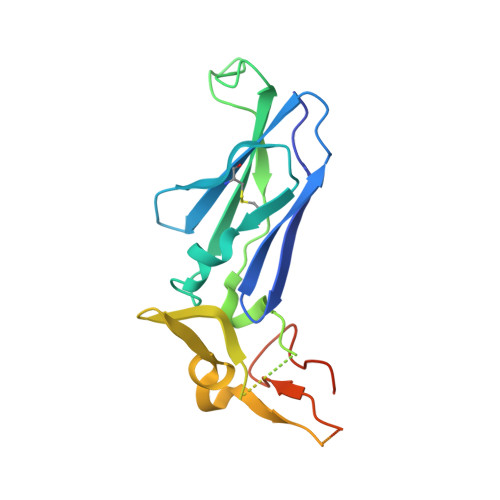
Discovery of novel heterocyclic factor VIIa inhibitors.
Rai, R., Kolesnikov, A., Sprengeler, P.A., Torkelson, S., Ton, T., Katz, B.A., Yu, C., Hendrix, J., Shrader, W.D., Stephens, R., Cabuslay, R., Sanford, E., Young, W.B.(2006) Bioorg Med Chem Lett 16: 2270-2273
- PubMed: 16460932 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2006.01.017
- Primary Citation Related Structures:
2F9B - PubMed Abstract:
Structure-activity relationships and binding mode of novel heterocyclic factor VIIa inhibitors will be described. In these inhibitors, a highly basic 5-amidinoindole moiety has been successfully replaced with a less basic 5-aminopyrrolo[3,2-b]pyridine scaffold.
- Celera, 180 Kimball Way, South San Francisco, CA 94080, USA. roopa.rai@hotmail.com
Organizational Affiliation: