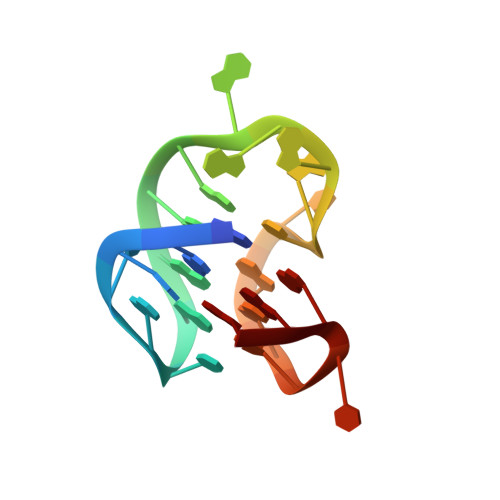
NMR solution structure of the major G-quadruplex structure formed in the human BCL2 promoter region.
Dai, J., Chen, D., Jones, R.A., Hurley, L.H., Yang, D.(2006) Nucleic Acids Res 34: 5133-5144
- PubMed: 16998187 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkl610
- Primary Citation Related Structures:
2F8U - PubMed Abstract:
BCL2 protein functions as an inhibitor of cell apoptosis and has been found to be aberrantly expressed in a wide range of human diseases. A highly GC-rich region upstream of the P1 promoter plays an important role in the transcriptional regulation of BCL2. Here we report the NMR solution structure of the major intramolecular G-quadruplex formed on the G-rich strand of this region in K+ solution. This well-defined mixed parallel/antiparallel-stranded G-quadruplex structure contains three G-tetrads of mixed G-arrangements, which are connected with two lateral loops and one side loop, and four grooves of different widths. The three loops interact with the core G-tetrads in a specific way that defines and stabilizes the overall G-quadruplex structure. The loop conformations are in accord with the experimental mutation and footprinting data. The first 3-nt loop adopts a lateral loop conformation and appears to determine the overall folding of the BCL2 G-quadruplex. The third 1-nt double-chain-reversal loop defines another example of a stable parallel-stranded structural motif using the G3NG3 sequence. Significantly, the distinct major BCL2 promoter G-quadruplex structure suggests that it can be specifically involved in gene modulation and can be an attractive target for pathway-specific drug design.
- College of Pharmacy, The University of Arizona, 1703 E. Mabel Street, Tucson, AZ 85721, USA.
Organizational Affiliation: