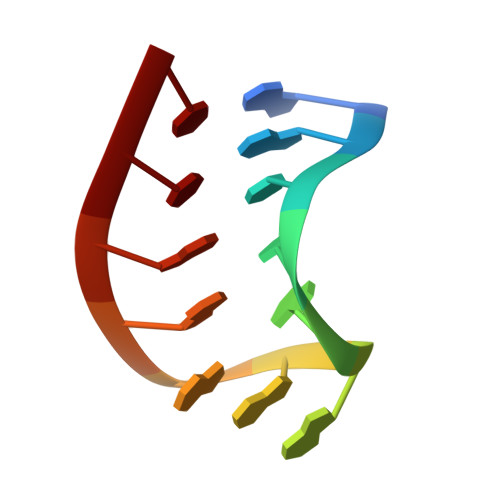
Solution structure of a GAAG tetraloop in helix 6 of SRP RNA from Pyrococcus furiosus
Okada, K., Takahashi, M., Sakamoto, T., Kawai, G., Nakamura, K., Kanai, A.(2006) Nucleosides Nucleotides Nucleic Acids 25: 383-395
- PubMed: 16838833 Search on PubMed
- DOI: https://doi.org/10.1080/15257770600683979
- Primary Citation Related Structures:
2F87 - PubMed Abstract:
The NMR structure of a 12-mer RNA derived from the helix 6 of SRP RNA from Pyrococcus furiosus, whose loop-closing base pair is U.G, was determined, and the structural and thermodynamic properties of the RNA were compared with those of a mutant RNA with the C:G closing base pair. Although the structures of the two RNAs are similar to each other and adopt the GNRR motif the conformational stabilities are significantly different to each other It was suggested that weaker stacking interaction of the GAAG loop with the U:G closing base pair in 12-mer RNA causes the lower conformational stability.
- Department of Life and Environmental Sciences, Faculty of Engineering, Chiba Institute of Technology, Chiba, Japan.
Organizational Affiliation: