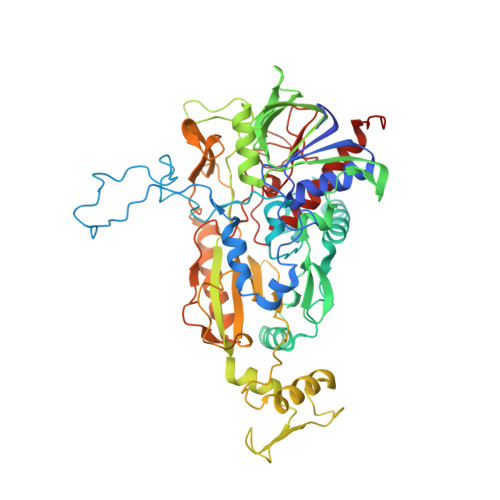
Reaction Geometry and Thermostable Variant of Pyranose 2-Oxidase from the White-Rot Fungus Peniophora sp.
Bannwarth, M., Heckmann-Pohl, D., Bastian, S., Giffhorn, F., Schulz, G.E.(2006) Biochemistry 45: 6587-6595
- PubMed: 16716069 Search on PubMed
- DOI: https://doi.org/10.1021/bi052465d
- Primary Citation Related Structures:
2F5V, 2F6C - PubMed Abstract:
Pyranose 2-oxidase catalyzes the oxidation of a number of carbohydrates using dioxygen; glucose, for example, is oxidized at carbon 2. The structure of pyranose 2-oxidase with the reaction product 2-keto-beta-d-glucose bound in the active center is reported in a new crystal form at 1.41 A resolution. The binding structure suggests that the alpha-anomer cannot be processed. The binding mode of the oxidized product was used to model other sugars accepted by the enzyme and to explain its specificity and catalytic rates. The reported structure at pH 6.0 shows a drastic conformational change in the loop of residues 454-461 (loop 454-461) at the active center compared to that of a closely homologous enzyme analyzed at pH 4.5 with a bound acetate inhibitor. In our structures, the loop is highly mobile and shifts to make way for the sugar to pass into the active center. Presumably, loop 454-461 functions as a gatekeeper. Apart from the wild-type enzyme, a thermostable variant was analyzed at 1.84 A resolution. In this variant, Glu542 is exchanged for a lysine. The observed stabilization could be a result of the mutated residue changing an ionic contact at a comparatively weak interface of the tetramer.
- Institut für Organische Chemie und Biochemie, Albert-Ludwigs-Universität, Albertstrasse 21, 79104 Freiburg im Breisgau, Germany.
Organizational Affiliation: