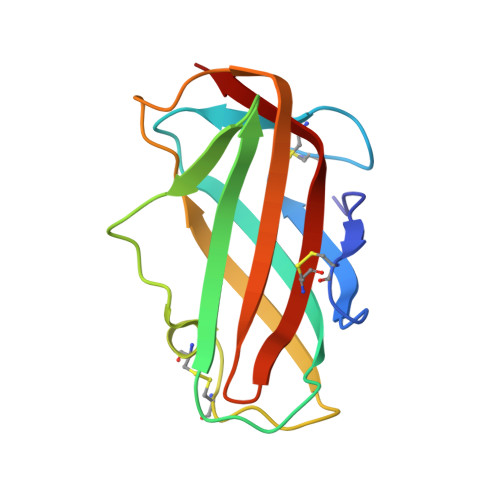
Crystal structure and some properties of a major house dust mite allergen, Derf 2
Suzuki, M., Tanaka, Y., Korematsu, S., Mikami, B., Minato, N.(2006) Biochem Biophys Res Commun 339: 679-686
- PubMed: 16313885 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2005.11.065
- Primary Citation Related Structures:
2F08 - PubMed Abstract:
Pyroglyphid house dust mites are a major source of allergens in house dust. Mite allergens sensitize and induce asthma, rhinitis, and eczema in a large portion of patients with allergic diseases. Here, the crystal structure of a major mite allergen, Derf 2, derived from Dermatophagoides farinae was solved by single isomorphous replacement method with anomalous scattering (SIRAS) at 2.1A resolution. The present study also demonstrated that the conformation of the allergen was critical in the determination of Th1/Th2 shift based on physicochemical and immunological analyses. This indicates that rigidly folded and singly dispersed structure is essentially required for the generation of Th2 type cells by the allergen, while conformational variant protein leads to Th1 skewing, irrespective of the same amino acid sequence. This structure/function relationship may allow us to develop a novel strategy for hyposensitization therapy in patients with allergic diseases triggered by house dust mite allergens.
- Laboratory of Immunology and Cell Biology, Graduate School of Biostudies, Kyoto University, Yoshidakonoe-Cho, Sakyo-Ku, Kyoto 606-8501, Japan.
Organizational Affiliation: