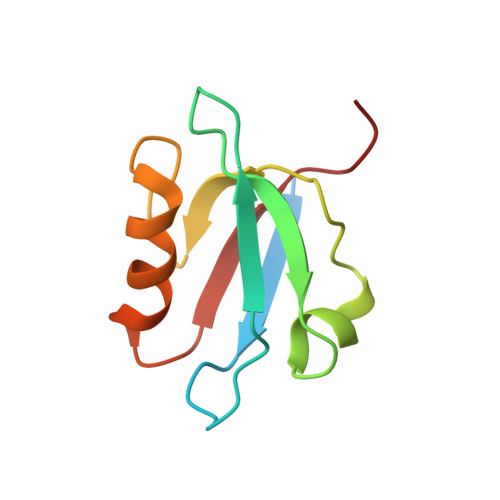
Solution structure of human erythroid p55 PDZ domain
Kusunoki, H., Kohno, T.(2006) Proteins 64: 804-807
- PubMed: 16741958 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21028
- Primary Citation Related Structures:
2EV8 - Mitsubishi Kagaku Institute of Life Sciences, Tokyo, Japan.
Organizational Affiliation: