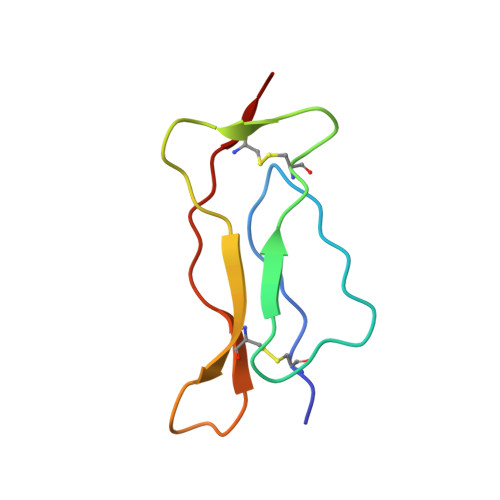
The structure of the interleukin-15 alpha receptor and its implications for ligand binding.
Lorenzen, I., Dingley, A.J., Jacques, Y., Grotzinger, J.(2006) J Biological Chem 281: 6642-6647
- PubMed: 16377614 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M513118200
- Primary Citation Related Structures:
2ERS - PubMed Abstract:
Interleukin (IL)-15 is a member of the small four alpha-helix bundle family of cytokines. IL-15 was discovered by its ability to mimic IL-2-mediated T-cell proliferation. Both cytokines share the beta and gamma receptor chains of the IL-2 receptor for signal transduction. However, in addition, they target specific alpha chain receptors IL-15Ralpha and IL-2Ralpha, respectively. The exceptionally high affinity binding of IL-15 to IL-15Ralpha is mediated by its sushi domain. Here we present the solution structure of the IL-15Ralpha sushi domain solved by NMR spectroscopy and a model of its complex with IL-15. The model shows that, rather than the familiar hydrophobic forces dominating the interaction interface between cytokines and their cognate receptors, the interaction between the IL-15 and IL-15Ralpha complex involves a large network of ionic interactions. This type of interaction explains the exceptionally high affinity of the IL-15.IL-15Ralpha complex, which is essential for the biological effects of this important cytokine and which is not observed in other cytokine/cytokine receptor complexes.
- Biochemisches Institut der Christian-Albrechts-Universität Kiel, Olshausenstrasse 40, 24118 Kiel, Germany.
Organizational Affiliation: