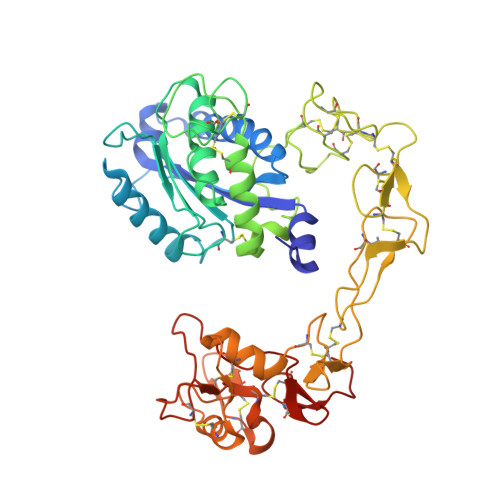
Crystal structures of VAP1 reveal ADAMs' MDC domain architecture and its unique C-shaped scaffold
Takeda, S., Igarashi, T., Mori, H., Araki, S.(2006) EMBO J 25: 2388-2396
- PubMed: 16688218
- DOI: https://doi.org/10.1038/sj.emboj.7601131
- Primary Citation Related Structures:
2ERO, 2ERP, 2ERQ - PubMed Abstract:
ADAMs (a disintegrin and metalloproteinase) are sheddases possessing extracellular metalloproteinase/disintegrin/cysteine-rich (MDC) domains. ADAMs uniquely display both proteolytic and adhesive activities on the cell surface, however, most of their physiological targets and adhesion mechanisms remain unclear. Here for the first time, we reveal the ADAMs' MDC architecture and a potential target-binding site by solving crystal structures of VAP1, a snake venom homolog of mammalian ADAMs. The D-domain protrudes from the M-domain opposing the catalytic site and constituting a C-shaped arm with cores of Ca2+ ions. The disintegrin-loop, supposed to interact with integrins, is packed by the C-domain and inaccessible for protein binding. Instead, the hyper-variable region (HVR) in the C-domain, which has a novel fold stabilized by the strictly conserved disulfide bridges, constitutes a potential protein-protein adhesive interface. The HVR is located at the distal end of the arm and faces toward the catalytic site. The C-shaped structure implies interplay between the ADAMs' proteolytic and adhesive domains and suggests a molecular mechanism for ADAMs' target recognition for shedding.
- Department of Cardiac Physiology, National Cardiovascular Center Research Institute, Suita, Osaka, Japan. stakeda@ri.ncvc.go.jp
Organizational Affiliation: