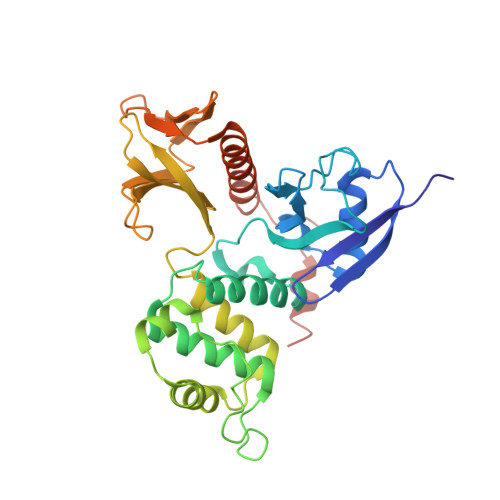
Structural basis of PSGL-1 binding to ERM proteins
Takai, Y., Kitano, K., Terawaki, S., Maesaki, R., Hakoshima, T.(2007) Genes Cells 12: 1329-1338
- PubMed: 18076570 Search on PubMed
- DOI: https://doi.org/10.1111/j.1365-2443.2007.01137.x
- Primary Citation Related Structures:
2EMT - PubMed Abstract:
P-selectin glycoprotein ligand-1 (PSGL-1), an adhesion molecule with O-glycosylated extracellular sialomucins, is involved in leukocyte inflammatory responses. On activation, ezrin-radixin-moesin (ERM) proteins mediate the redistribution of PSGL-1 on polarized cell surfaces to facilitate binding to target molecules. ERM proteins recognize a short binding motif, Motif-1, conserved in cytoplasmic tails of adhesion molecules, whereas PSGL-1 lacks Motif-1 residues important for binding to ERM proteins. The crystal structure of the complex between the radixin FERM domain and a PSGL-1 juxtamembrane peptide reveals that the peptide binds the groove of FERM subdomain C by forming a beta-strand associated with strand beta5C, followed by a loop flipped out towards the solvent. The Motif-1 3(10) helix present in the FERM-ICAM-2 complex is absent in PSGL-1 given the absence of a critical Motif-1 alanine residue, and PSGL-1 reduces its contact area with subdomain C. Non-conserved positions are occupied by large residues Met9 and His8, which stabilize peptide conformation and enhance groove binding. Non-conserved residues play an important role in compensating for loss of binding energy resulting from the absence of conserved residues important for binding.
- Structural Biology Laboratory, Nara Institute of Science and Technology, Nara, Japan.
Organizational Affiliation: