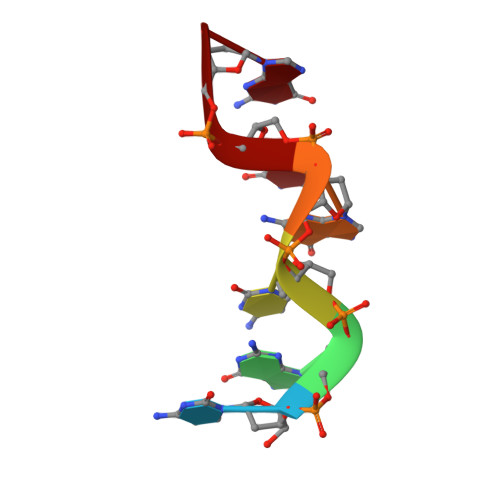
The rare crystallographic structure of d(CGCGCG)(2): The natural spermidine molecule bound to the minor groove of left-handed Z-DNA d(CGCGCG)(2) at 10 degrees C
Ohishi, H., Tozuka, Y., Da-Yang, Z., Ishida, T., Nakatani, K.(2007) Biochem Biophys Res Commun 358: 24-28
- PubMed: 17467661 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2007.04.026
- Primary Citation Related Structures:
2ELG - PubMed Abstract:
Several crystal structure analyses of complexes of synthetic polyamine compounds, including N(1)-(2-(2-aminoethylamino))ethyl)ethane-1,2-diamine PA(222) and N(1)-(2-(2-(2-aminoethylamino)ethylamino)ethyl)ethane-1,2-diamine PA(2222), and left-handed Z-DNA d(CGCGCG)(2) have been reported. However, until now, there have been no examples of naturally occurring polyamines bound to the minor groove of the left-handed Z-DNA of d(CGCGCG)(2) molecule. We have found that spermidine, a natural polyamine, is connected to the minor groove of left-handed Z-DNA of d(CGCGCG)(2) molecule in a crystalline complex grown at 10 degrees C. The electron density of the DNA molecule was clear enough to determine that the spermidine was connected in the minor groove of two symmetry related molecules of left-handed Z-DNA d(CGCGCG)(2). This is the first example that a spermidine molecule can form a bridge conformation between two symmetry related molecules of left-handed Z-DNA d(CGCGCG)(2) in the minor groove.
- Osaka University of Pharmaceutical Sciences, 4-20-1, Nasahara, Takatsuki, Osaka 569-1094, Japan. ohishi@gly.oups.ac.jp
Organizational Affiliation: