Structure of a SUMO-binding-motif Mimic Bound to Smt3p-Ubc9p: Conservation of a Non-covalent Ubiquitin-like Protein-E2 Complex as a Platform for Selective Interactions within a SUMO Pathway
Duda, D.M., van Waardenburg, R.C.A.M., Borg, L.A., McGarity, S., Nourse, A., Waddell, M.B., Bjornsti, M.A., Schulman, B.A.(2007) J Mol Biology 369: 619-630
- PubMed: 17475278
- DOI: https://doi.org/10.1016/j.jmb.2007.04.007
- Primary Citation Related Structures:
2EKE - PubMed Abstract:
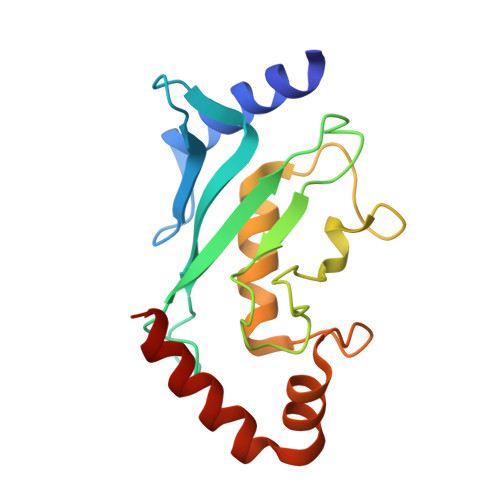
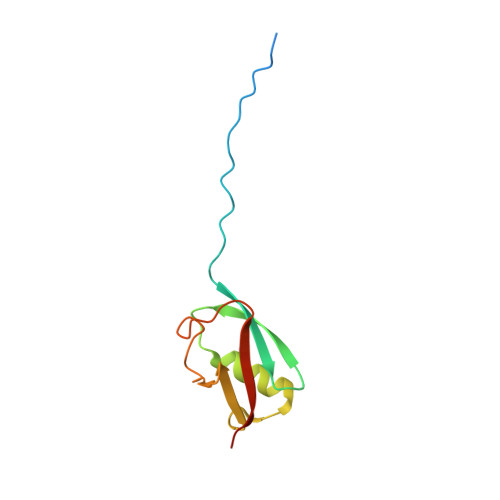
The SUMO ubiquitin-like proteins play regulatory roles in cell division, transcription, DNA repair, and protein subcellular localization. Paralleling other ubiquitin-like proteins, SUMO proteins are proteolytically processed to maturity, conjugated to targets by E1-E2-E3 cascades, and subsequently recognized by specific downstream effectors containing a SUMO-binding motif (SBM). SUMO and its E2 from the budding yeast Saccharomyces cerevisiae, Smt3p and Ubc9p, are encoded by essential genes. Here we describe the 1.9 A resolution crystal structure of a non-covalent Smt3p-Ubc9p complex. Unexpectedly, a heterologous portion of the crystallized complex derived from the expression construct mimics an SBM, and binds Smt3p in a manner resembling SBM binding to human SUMO family members. In the complex, Smt3p binds a surface distal from Ubc9's catalytic cysteine. The structure implies that a single molecule of Smt3p cannot bind concurrently to both the non-covalent binding site and the catalytic cysteine of a single Ubc9p molecule. However, formation of higher-order complexes can occur, where a single Smt3p covalently linked to one Ubc9p's catalytic cysteine also binds non-covalently to another molecule of Ubc9p. Comparison with other structures from the SUMO pathway suggests that formation of the non-covalent Smt3p-Ubc9p complex occurs mutually exclusively with many other Smt3p and Ubc9p interactions in the conjugation cascade. By contrast, high-resolution insights into how Smt3p-Ubc9p can also interact with downstream recognition machineries come from contacts with the SBM mimic. Interestingly, the overall architecture of the Smt3p-Ubc9p complex is strikingly similar to recent structures from the ubiquitin pathway. The results imply that non-covalent ubiquitin-like protein-E2 complexes are conserved platforms, which function as parts of larger assemblies involved in many protein post-translational regulatory pathways.
- Department of Structural Biology and Genetics, St. Jude Children's Research Hospital, Memphis, TN 38105, USA.
Organizational Affiliation: