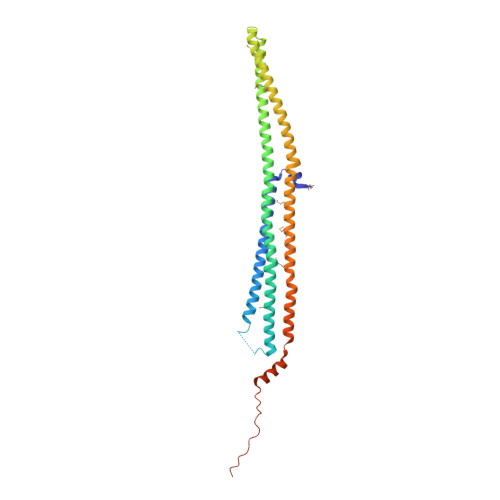
Curved EFC/F-BAR-Domain Dimers Are Joined End to End into a Filament for Membrane Invagination in Endocytosis
Shimada, A., Niwa, H., Tsujita, K., Suetsugu, S., Nitta, K., Hanawa-Suetsugu, K., Akasaka, R., Nishino, Y., Toyama, M., Chen, L., Liu, Z.-J., Wang, B.-C., Yamamoto, M., Terada, T., Miyazawa, A., Tanaka, A., Sugano, S., Shirouzu, M., Nagayama, K., Takenawa, T., Yokoyama, S.(2007) Cell 129: 761-772
- PubMed: 17512409 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2007.03.040
- Primary Citation Related Structures:
2EFK, 2EFL - PubMed Abstract:
Pombe Cdc15 homology (PCH) proteins play an important role in a variety of actin-based processes, including clathrin-mediated endocytosis (CME). The defining feature of the PCH proteins is an evolutionarily conserved EFC/F-BAR domain for membrane association and tubulation. In the present study, we solved the crystal structures of the EFC domains of human FBP17 and CIP4. The structures revealed a gently curved helical-bundle dimer of approximately 220 A in length, which forms filaments through end-to-end interactions in the crystals. The curved EFC dimer fits a tubular membrane with an approximately 600 A diameter. We subsequently proposed a model in which the curved EFC filament drives tubulation. In fact, striation of tubular membranes was observed by phase-contrast cryo-transmission electron microscopy, and mutations that impaired filament formation also impaired membrane tubulation and cell membrane invagination. Furthermore, FBP17 is recruited to clathrin-coated pits in the late stage of CME, indicating its physiological role.
- RIKEN SPring-8 Center, Harima Institute, 1-1-1 Kouto, Sayo, Hyogo 679-5148, Japan.
Organizational Affiliation: