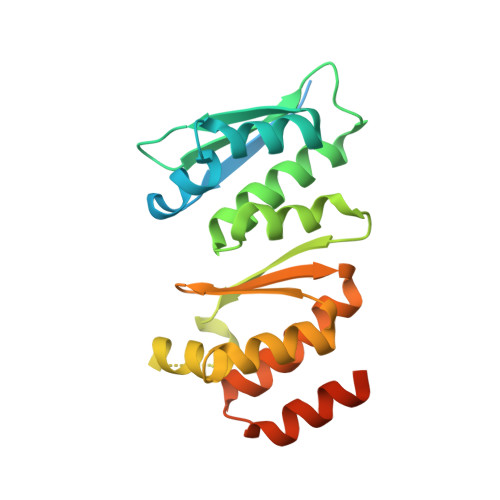
Crystal structure of Dim2p: a preribosomal RNA processing factor, from Pyrococcus horikoshii OT3 at 2.30 A
Jia, M.Z., Ohtsuka, J., Lee, W.C., Nagata, K., Tanokura, M.(2007) Proteins 69: 428-432
- PubMed: 17654551 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21381
- Primary Citation Related Structures:
2E3U - Department of Applied Biological Chemistry, Graduate School of Agricultural and Life Sciences, University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan.
Organizational Affiliation: