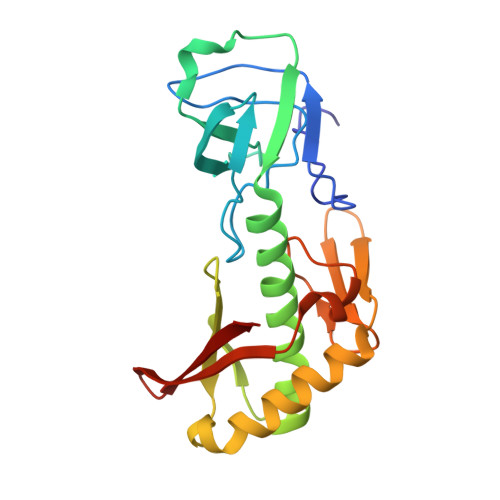
Structure of the AlaX-M trans-editing enzyme from Pyrococcus horikoshii
Fukunaga, R., Yokoyama, S.(2007) Acta Crystallogr D Biol Crystallogr 63: 390-400
- PubMed: 17327676
- DOI: https://doi.org/10.1107/S090744490605640X
- Primary Citation Related Structures:
2E1B - PubMed Abstract:
The editing domain of alanyl-tRNA synthetase (AlaRS) contributes to high-fidelity aminoacylation by hydrolyzing (editing) the incorrect products Ser-tRNA(Ala) and Gly-tRNA(Ala) (cis-editing). The AlaX protein shares sequence homology to the editing domain of AlaRS. There are three types of AlaX proteins, with different numbers of amino-acid residues (AlaX-S, AlaX-M and AlaX-L). In this report, AlaX-M from Pyrococcus horikoshii is shown to deacylate Ser-tRNA(Ala) and Gly-tRNA(Ala) (trans-editing). The crystal structure of P. horikoshii AlaX-M has been determined at 2.7 A resolution. AlaX-M consists of an N-terminal domain (N-domain) and a C-terminal domain (C-domain). A zinc ion is coordinated by the conserved zinc-binding cluster in the C-domain, which is expected to be the enzymatic active site. The glycine-rich motif, consisting of successive conserved glycine residues in the N-domain, forms a loop (the 'glycine-rich loop'). The glycine-rich loop is located near the active site and may be involved in substrate recognition and/or catalysis.
- Department of Biophysics and Biochemistry, Graduate School of Science, University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-0033, Japan.
Organizational Affiliation: