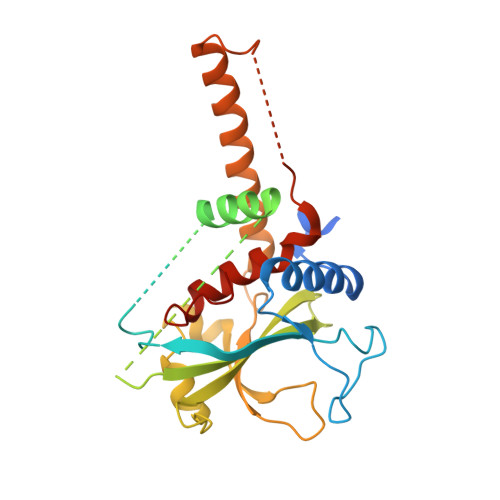
The crystal structure of Atg3, an autophagy-related ubiquitin carrier protein (E2) enzyme that mediates Atg8 lipidation
Yamada, Y., Suzuki, N.N., Hanada, T., Ichimura, Y., Kumeta, H., Fujioka, Y., Ohsumi, Y., Inagaki, F.(2007) J Biological Chem 282: 8036-8043
- PubMed: 17227760 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M611473200
- Primary Citation Related Structures:
2DYT - PubMed Abstract:
Atg3 is an E2-like enzyme that catalyzes the conjugation of Atg8 and phosphatidylethanolamine (PE). The Atg8-PE conjugate is essential for autophagy, which is the bulk degradation process of cytoplasmic components by the vacuolar/lysosomal system. We report here the crystal structure of Saccharomyces cerevisiae Atg3 at 2.5-A resolution. Atg3 has an alpha/beta-fold, and its core region is topologically similar to canonical E2 enzymes. Atg3 has two regions inserted in the core region, one of which consists of approximately 80 residues and has a random coil structure in solution and another with a long alpha-helical structure that protrudes from the core region as far as 30 A. In vivo and in vitro analyses suggested that the former region is responsible for binding Atg7, an E1-like enzyme, and that the latter is responsible for binding Atg8. A sulfate ion was bound near the catalytic cysteine of Atg3, suggesting a possible binding site for the phosphate moiety of PE. The structure of Atg3 provides a molecular basis for understanding the unique lipidation reaction that Atg3 carries out.
- Department of Structural Biology, Graduate School of Pharmaceutical Sciences, Hokkaido University, N-12, W-6, Kita-ku, Sapporo 060-0812, Japan.
Organizational Affiliation: