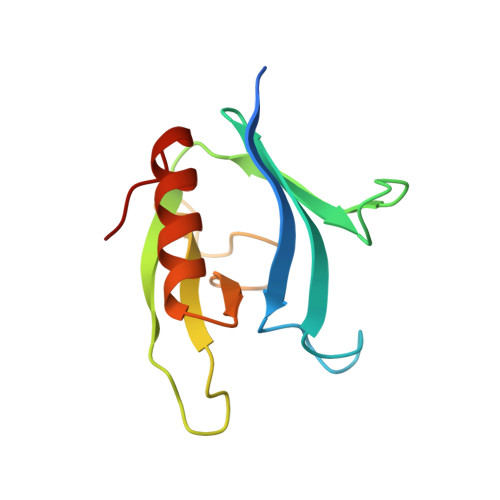
Crystal structure of the pleckstrin homology domain from dynamin.
Timm, D., Salim, K., Gout, I., Guruprasad, L., Waterfield, M., Blundell, T.(1994) Nat Struct Biol 1: 782-788
- PubMed: 7634088 Search on PubMed
- DOI: https://doi.org/10.1038/nsb1194-782
- Primary Citation Related Structures:
2DYN - PubMed Abstract:
The pleckstrin homology (PH) domain is a conserved module present in many signal transducing and cytoskeletal proteins. Here we report the 2.8 A crystal structure of the PH domain from dynamin. This domain consists of seven beta-strands forming two roughly orthogonal antiparallel beta-sheets terminating with an amphipathic alpha-helix. The structure also reveals a non-covalent dimeric association of the PH domain and a hydrophobic pocket surrounded by a charged rim. The dynamin PH domain structure is discussed in relation to its potential role in mediating interactions between proteins.
- Department of Crystallography, Birkbeck College, London, UK.
Organizational Affiliation: