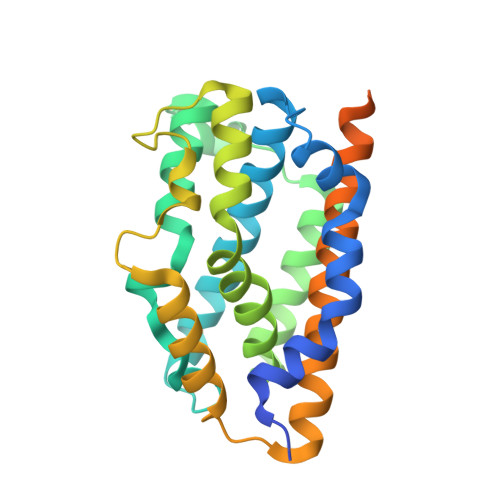
X-ray crystallographic and biochemical characterization of the inhibitory action of an imidazole-dioxolane compound on heme oxygenase
Sugishima, M., Higashimoto, Y., Oishi, T., Takahashi, H., Sakamoto, H., Noguchi, M., Fukuyama, K.(2007) Biochemistry 46: 1860-1867
- PubMed: 17253780 Search on PubMed
- DOI: https://doi.org/10.1021/bi062264p
- Primary Citation Related Structures:
2DY5 - PubMed Abstract:
Heme oxygenase (HO) catalyzes the regiospecific cleavage of the porphyrin ring of heme using reducing equivalents and O2 to produce biliverdin, iron, and CO. Because CO has a cytoprotective effect through the p38-MAPK pathway, HO is a potential therapeutic target in cancer. In fact, inhibition of the HO isoform HO-1 reduces Kaposi sarcoma tumor growth. Imidazole-dioxolane compounds have recently attracted attention because they have been reported to specifically inhibit HO-1, but not HO-2, unlike Cr-containing protoporphyrin IX, a classical inhibitor of HO, that inhibits not only both HO isoforms but also other hemoproteins. The inhibitory mechanism of imidazole-dioxolane compounds, however, has not yet been characterized. Here, we determine the crystal structure of the ternary complex of rat HO-1, heme, and an imidazole-dioxolane compound, 2-[2-(4-chlorophenyl)ethyl]-2-[(1H-imidazol-1-yl)methyl]-1,3-dioxolane. This compound bound on the distal side of the heme iron, where the imidazole and 4-chlorophenyl groups were bound to the heme iron and the hydrophobic cavity in HO, respectively. Binding of the bulky inhibitor in the narrow distal pocket shifted the distal helix to open the distal site and moved both the heme and the proximal helix. Furthermore, the biochemical characterization revealed that the catalytic reactions of both HO-1 and HO-2 were completely stopped after the formation of verdoheme in the presence of the imidazole-dioxolane compound. This result should be mainly due to the lower reactivity of the inhibitor-bound verdoheme with O2 compared to the reactivity of the inhibitor-bound heme with O2.
- Department of Medical Biochemistry, Kurume University School of Medicine, Kurume, Fukuoka 830-0011, Japan.
Organizational Affiliation: