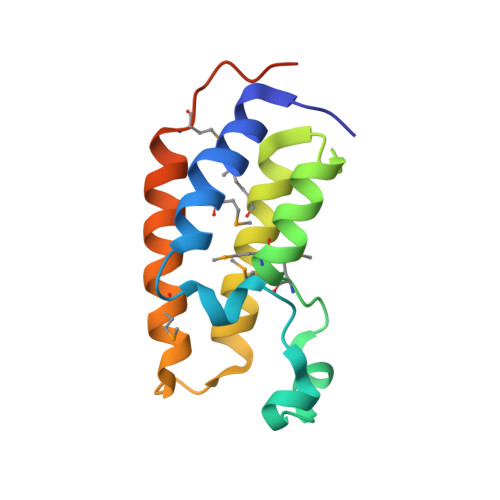
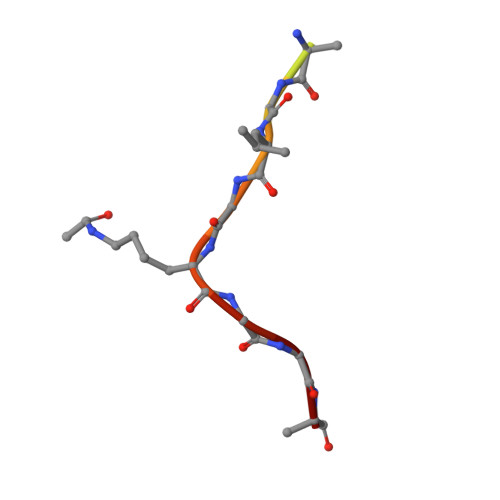
Structural Basis for Acetylated Histone H4 Recognition by the Human BRD2 Bromodomain.
Umehara, T., Nakamura, Y., Jang, M.K., Nakano, K., Tanaka, A., Ozato, K., Padmanabhan, B., Yokoyama, S.(2010) J Biological Chem 285: 7610-7618
- PubMed: 20048151
- DOI: https://doi.org/10.1074/jbc.M109.062422
- Primary Citation Related Structures:
2DVQ, 2DVR, 2DVS - PubMed Abstract:
Recognition of acetylated chromatin by the bromodomains and extra-terminal domain (BET) family proteins is a hallmark for transcriptional activation and anchoring viral genomes to mitotic chromosomes of the host. One of the BET family proteins BRD2 interacts with acetylated chromatin during mitosis and leads to transcriptional activation in culture cells. Here, we report the crystal structures of the N-terminal bromodomain of human BRD2 (BRD2-BD1; residues 74-194) in complex with each of three different Lys-12-acetylated H4 peptides. The BRD2-BD1 recognizes the H4 tail acetylated at Lys-12 (H4K12ac), whereas the side chain of hypoacetylated Lys-8 of H4 binds at the cavity of the dimer interface of BRD2-BD1. From binding studies, we identified the BRD2-BD1 residues that are responsible for recognition of the Lys-12-acetylated H4 tail. In addition, mutation to Lys-8 in the Lys-12-acetylated H4 tail decreased the binding to BRD2-BD1, implicating the critical role of Lys-8 in the Lys-12-acetylated H4 tail for the recognition by BRD2-BD1. Our findings provide a structural basis for deciphering the histone code by the BET bromodomain through the binding with a long segment of the histone H4 tail, which presumably prevents erasure of the histone code during the cell cycle.
- RIKEN Systems and Structural Biology Center, 1-7-22 Suehiro-cho, Tsurumi, Yokohama 230-0045, Japan.
Organizational Affiliation: