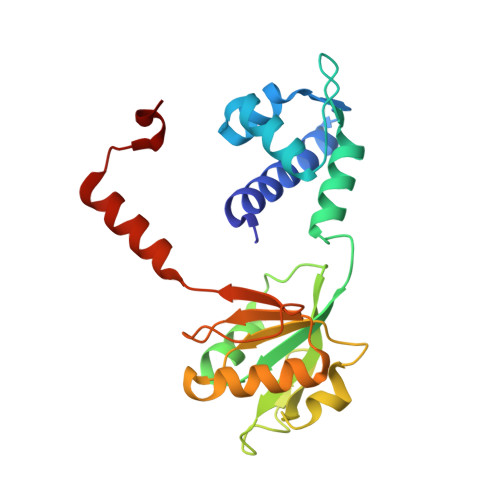
Crystal structure of TTHA1657 (AT-rich DNA-binding protein; p25) from Thermus thermophilus HB8 at 2.16 A resolution
Nakamura, A., Sosa, A., Komori, H., Kita, A., Miki, K.(2007) Proteins 66: 755-759
- PubMed: 17154156 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21222
- Primary Citation Related Structures:
2DT5 - Department of Chemistry, Graduate School of Science, Kyoto University, Sakyo-ku, Kyoto 606-8502, Japan.
Organizational Affiliation: