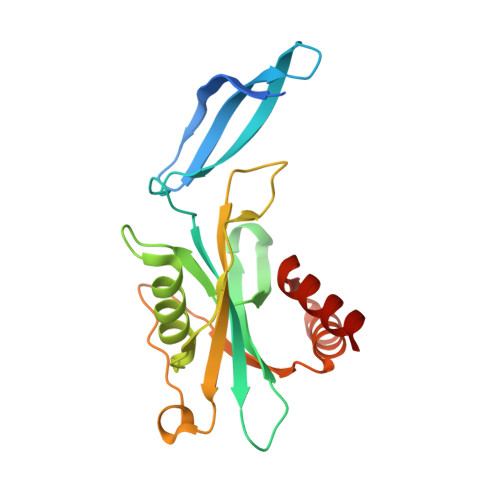
Crystal Structures of Human NUDT5 Reveal Insights into the Structural Basis of the Substrate Specificity
Zha, M., Zhong, C., Peng, Y., Hu, H., Ding, J.(2006) J Mol Biology 364: 1021-1033
- PubMed: 17052728 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.09.078
- Primary Citation Related Structures:
2DSB, 2DSC, 2DSD - PubMed Abstract:
Human NUDT5 (hNUDT5) is an ADP-ribose pyrophosphatase (ADPRase) belonging to the Nudix hydrolase superfamily. It presumably plays important roles in controlling the intracellular level of ADP-ribose (ADPR) to prevent non-enzymatic ADP-ribosylation by hydrolyzing ADPR to AMP and ribose 5'-phosphate. We report here the crystal structures of hNUDT5 in apo form, in complex with ADPR, and in complex with AMP with bound Mg2+. hNUDT5 forms a homodimer with substantial domain swapping and assumes a structure more similar to Escherichia coli ADPRase ORF209 than human ADPRase NUDT9. The adenine moiety of the substrates is specifically recognized by the enzyme via hydrogen-bonding interactions between N1 and N6 of the base and Glu47 of one subunit, and between N7 of the base and Arg51 of the other subunit, providing the molecular basis for the high selectivity of hNUDT5 for ADP-sugars over other sugar nucleotides. Structural comparisons with E. coli ADPRase ORF209 and ADPXase ORF186 indicate that the existence of an aromatic residue on loop L8 in ORF186 seems to be positively correlated with its enzymatic activity on APnA, whereas hNUDT5 and ORF209 contain no such residue and thus have low or no activities on APnA.
- State Key Laboratory of Molecular Biology, Institute of Biochemistry and Cell Biology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, Shanghai 200031, China.
Organizational Affiliation: