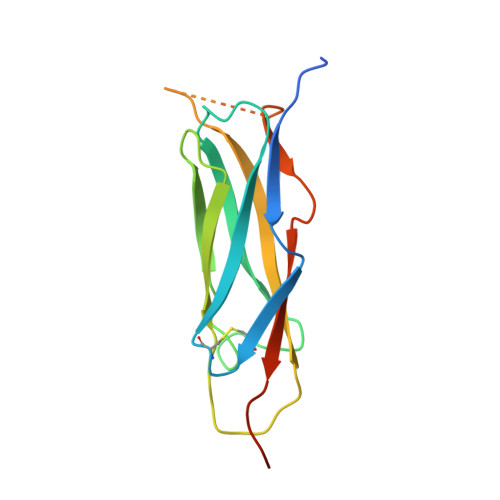
The crystal structure of the primary Ca2+ sensor of the na+/ca2+ exchanger reveals a novel Ca2+ binding motif.
Nicoll, D.A., Sawaya, M., Kwon, S., Cascio, D., Philipson, K.D., Abramson, J.(2006) J Biological Chem 281: 21577-21581
- PubMed: 16774926 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.C600117200
- Primary Citation Related Structures:
2DPK - PubMed Abstract:
The Na+/Ca2+ exchanger is a plasma membrane protein that regulates intracellular Ca2+ levels in cardiac myocytes. Transport activity is governed by Ca2+, and the primary Ca2+ sensor (CBD1) is located in a large cytoplasmic loop connecting two transmembrane helices. The binding of Ca2+ to the CBD1 sensory domain results in conformational changes that stimulate the exchanger to extrude Ca2+. Here, we present a crystal structure of CBD1 at 2.5A resolution, which reveals a novel Ca2+ binding site consisting of four Ca2+ ions arranged in a tight planar cluster. This intricate coordination pattern for a Ca2+ binding cluster is indicative of a highly sensitive Ca2+ sensor and may represent a general platform for Ca2+ sensing.
- Department of Physiology and the Cardiovascular Research Laboratories, David Geffen School of Medicine, University of California, Los Angeles, California 90095.
Organizational Affiliation: