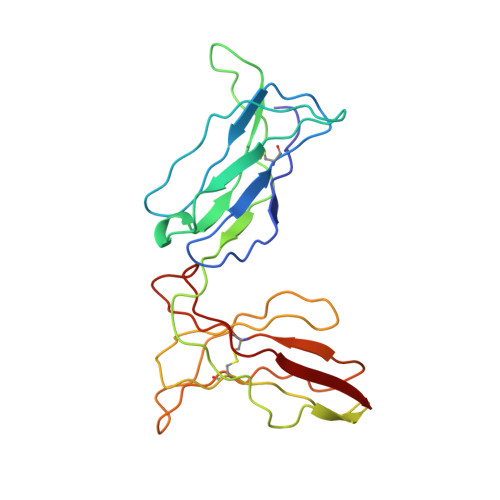
Crystal structure of the HLA-Cw3 allotype-specific killer cell inhibitory receptor KIR2DL2
Snyder, G.A., Brooks, A.G., Sun, P.D.(1999) Proc Natl Acad Sci U S A 96: 3864-3869
- PubMed: 10097129 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.96.7.3864
- Primary Citation Related Structures:
2DL2, 2DLI - PubMed Abstract:
Killer cell inhibitory receptors (KIR) protect class I HLAs expressing target cells from natural killer (NK) cell-mediated lysis. To understand the molecular basis of this receptor-ligand recognition, we have crystallized the extracellular ligand-binding domains of KIR2DL2, a member of the Ig superfamily receptors that recognize HLA-Cw1, 3, 7, and 8 allotypes. The structure was determined in two different crystal forms, an orthorhombic P212121 and a trigonal P3221 space group, to resolutions of 3.0 and 2.9 A, respectively. The overall fold of this structure, like KIR2DL1, exhibits K-type Ig topology with cis-proline residues in both domains that define beta-strand switching, which sets KIR apart from the C2-type hematopoietic growth hormone receptor fold. The hinge angle of KIR2DL2 is approximately 80 degrees, 14 degrees larger than that observed in KIR2DL1 despite the existence of conserved hydrophobic residues near the hinge region. There is also a 5 degrees difference in the observed hinge angles in two crystal forms of 2DL2, suggesting that the interdomain hinge angle is not fixed. The putative ligand-binding site is formed by residues from several variable loops with charge distribution apparently complementary to that of HLA-C. The packing of the receptors in the orthorhombic crystal form offers an intriguing model for receptor aggregation on the cell surface.
- Department of Biochemistry, Molecular Biology, and Cell Biology, 2153 Sheridan, O. T. Hogan Hall, Room 2-100, Northwestern University, Evanston, IL 60208, USA.
Organizational Affiliation: